Phragmoporthe conformis (Berk. & Broome) Petr., Annls mycol. 39(4/6): 285 (1941)
Saprobic on Alnus glutinosa L. Sexual morph Appearing as conical, pustules on the host surface. Ascomata perithecial, minutely stromatic, immersed, erumpent. Perithecia 700–770μm diam. (n =20), solitary, immersed in or directly below the host epidermis, globose, membranous, dark brown to black, with a periphysate ostiole. Peridium 14–38 μm (x = 22 μm, n = 10) wide, comprising 7–15 cell layers, outer layers heavily pigmented, thin-walled, comprising dark brown cells of textura angularis, inner layers composed of hyaline to brown, thin-walled, flat cells of textura angularis. Hamathecium lacking paraphyses. Asci 60–80 × 17–24 μm (x = 72×19.5 μm, n = 30), 8-spored, unitunicate, clavate, straight, short pedicellate, apically rounded or truncate, with a refractive, J- apical ring. Ascospores 19–24 × 6.5– 8 μm (x = 22 × 7 μm, n = 50), multi-seriate, fusiform, mainly with 3 transverse septa, occasionally constricted at septum, hyaline, smooth and thick-walled, without a mucilaginous sheath or appendages. Asexual morph Undetermined
Culture characteristics: Colonies growing on MEA, slow growing, reaching 4 cm diam. in 21d at 16 °C on MEA, white, dense, moderate aerial mycelium on the surface, underneath similar in colour, margins even.
Material examined: ITALY, Forlì-Cesena Province, Lago Pontini-Bagno di Romagna, dead branches of Alnus glutinosa (L.) Gaertn. (Betulaceae), 26 May 2014, Erio Camporesi, IT 1892 (MFLU 15–2662 reference specimen designated here), also in HKAS 92498, living culture, MFLUCC 14–0567.
Notes: The putatively named strain of Phragmoporthe conformis (CBS 109793) clustered with our newly collected
strain (MFLU 14–0567), collected fromItaly, on a dead a stem of Alnus glutinosa. Berkeley and Broome (1852) originally described Phragmoporthe conformis as Sphaeria conformis on Alnus spp. from the UK. Later Petrak (1941) synonymized Sphaeria conformis under Phragmoporthe conformis. The ascomata, size of asci and ascospores of our strain are typical of P. conformis (Petrak 1941) and the molecular data is identical to CBS 109793. We therefore designate our collection as a reference specimen of P. conformis to stabilize the taxonomy of the genus.
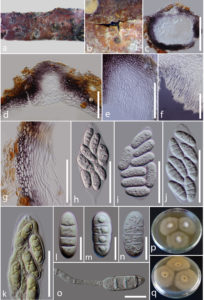
Fig. 1 Maximum Likelihood tree resulting from analysis of combined LSU, ITS and TEF-1α sequence data for taxa of the family Gnomoniaceae. Maximum likelihood bootstrap support values greater than 50 % are shown near the nodes. New taxa are in blue and ex-type strains in bold. The tree is rooted with Valsella salicis and Leucostoma niveum.
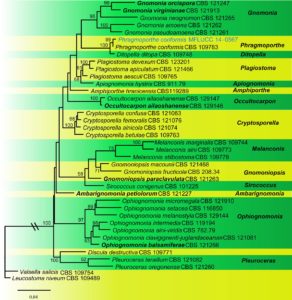
Fig. 2 Phragmoporthe conformis (MFLU 15–2662, reference specimen) a, b Appearance of ascomata on host substrate c Section of ascoma d Transverse section through ostiole e, f Periphyses g Close up of peridium h–j Asci k Close up of apical ascus strained inMelzer’s reagent l–n Ascospores o Germinating spore p, q Colonies growing on MEA. Scale bars: c = 500 μm, d, e = 100 μm, f–j = 50 μm, k = 20 μm, l– o = 10 μm