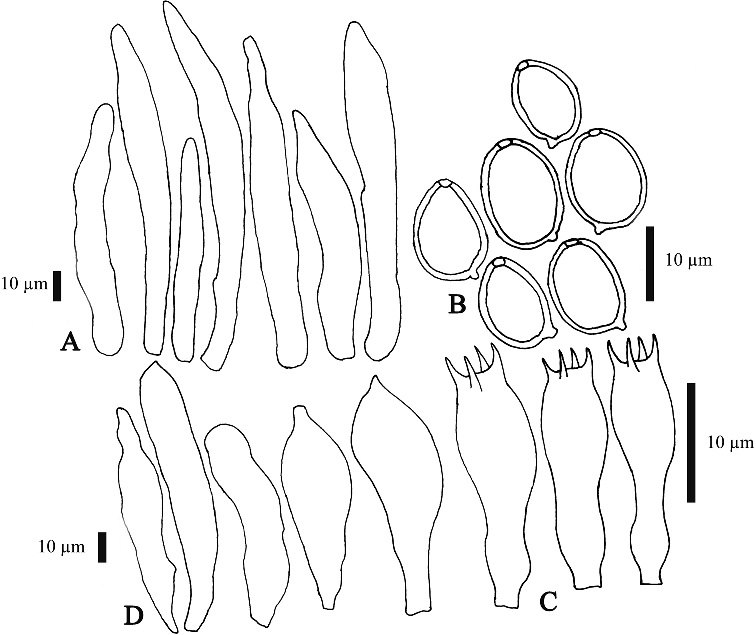
Leucocoprinus cretaceus (Bull.) Locq., Bull. mens. Soc. linn. Soc. Bot. Lyon 14: 93 (1945)
Facesoffungi number: FoF 3137
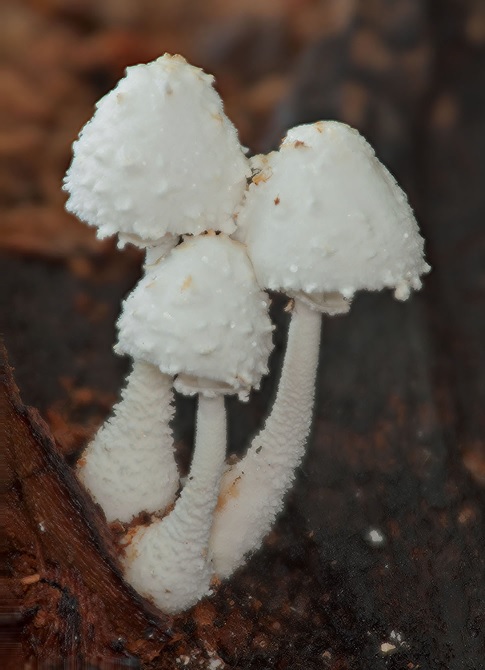
Pileus 3–6 cm broad, hemispherical when young, expanding to planoconvex to campanulate, margin decurved, slightly sulcate-striate, when young with many, dense, soft floccules, readily collapsing or wearing off to leave farinose covering; light buff to white at center, white elsewhere. Lamellae distantly free, rather broad, thin; white, edge slightly fimbriate. Stipe 5–8 cm long, 4–6 mm broad at apex, gradually enlarged downward to a broadly clavate to somewhat fusiform 6–14 mm broad base; sometimes coarsely farinose to slightly flocculose-farinose below annulus, subfarinose above; white to ivory yellow tinted. Annulus white, very soft, rather flaring, median to superior. Basidiospores 8–11 × 5–8 μm, Q=1.32–1.47, metachromatic, dextrinoid, ellipsoid to amygdaliform, thick-walled, with an apical germ pore covered with a hyaline lens. Basidia 16–22 × 9–12 μm, pyriform to clavate with a bulbous base, 4-spored, surrounded by 4 pseudoparaphyses. Sterigmata 1–2 × 0.7–1.5 μm. Pseudoparaphyses 10–13 × 7–10 μm, sphaeropedunculate to broadly clavate to pyriform, very broad point of attachment. Cheilocystidia 27–75 × 7–18 μm, thin-walled, hyaline, subcylindrical to narrowly fusiform to slightly narrowly lageniform, mucronate, rarely obtuse, moderately pedicellate. Pileus covering a sparse layer of terminal cells essentially absent on mature pilei, most prevalent near the center of young pilei, 45–117 × 7–9 μm, cylindrical to narrowly lageniform, with flexuous necks. Stipe covering structure like that of pileus, with terminal cells even more sparse. Stipitipellis a cutis of narrowly cylindrical, 5–8 μm broad elements. Clamp connections absent.
Habit, habitat and distribution: as a group, usually on manure or wood chips, May to August. Our collection was collected on cow manure at Hanthana Mountains Range, Peradeniya.
Specimen examined: SRI LANKA, Kandy District, Hanthana Mountains Range, 6 June 2012, Samantha C. Karunarathna (MFLU 12-1893, new record). GenBank number ITS:KY649466.
Notes: This is widely distributed in North America and this is the first report of L. cretaceus with molecular phylogenetic confirmation from Sri Lanka.
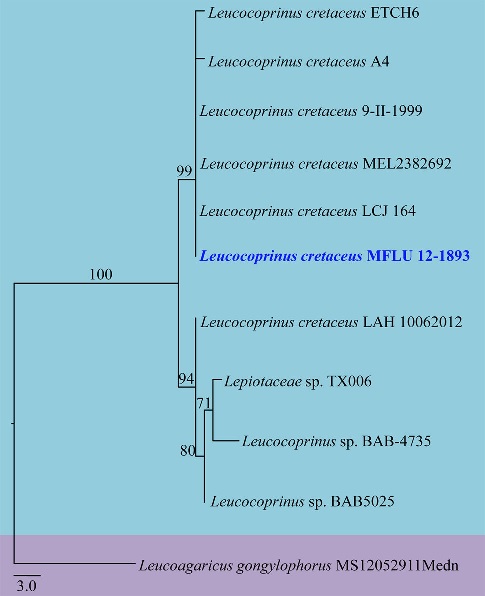
Phylogenetic relationships inferred from maximum parsimony analysis of ITS-rDNA sequences of 11 taxa. The percentages of replicate trees in which the associated taxa clustered together in the bootstrap test (1,000 replicates) are shown next to the branches. The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. The new Sri Lankan record: Leucocoprinus cretaceus having GenBank accession number KY649466 and herbarium number MFLU 12-1893 (ITS-rDNA) is shown in bold and blue. The topology is rooted with Leucoagaricus gongylophorus. Evolutionary analysis was conducted in PAUP 4.0b 10.