Sordariomycetidae O.E. Erikss. & Winka, Myconet 1(1): 10 (1997)
Index Fungorum number: IF 90351; MycoBank number: MB 90351; Facesoffungi number: FoF 06513;
The subclass Sordariomycetidae was established by Eriksson & Winka (1997) and comprised six orders, 12 families and two families incertae sedis. Members of this subclass are mainly characterized by dark ascomata with inoperculate, unitunicate asci and occur in terrestrial, aquatic and marine habitats and are widely distributed as plant and animal pathogens, endophytes, saprobes as well as coprophilous and lichenicolous taxa (Maharachchikumbura et al. 2015, 2016b, Huang et al. 2019). They are mycophilic, rich in coprophilous taxa and associated with invertebrates, and their ecological aspects and biotechnological potential have been researched (Zhang et al. 2006, Raghukumar 2008, Bovio et al. 2018). An MCC tree based on a combined SSU, LSU, tef1 and rpb2 sequence data revealed that this subclass evolved around 145–216 MYA (Hongsanan et al. 2017, Hyde et al. 2017a). The divergence time for Sordariomycetidae is estimated as 247 MYA (Fig. 2). Currently, there are eight orders and 19 families in this subclass (this paper).
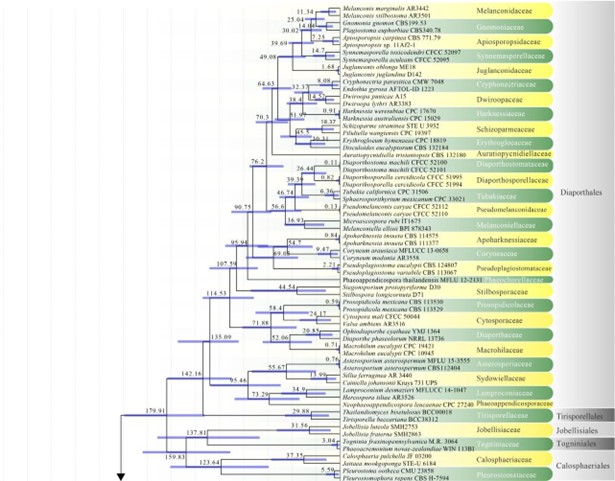
Figure 2 – The maximum clade credibility (MCC) tree, using the same dataset from Fig. 1. This analysis was performed in BEAST v1.10.2. The crown age of Sordariomycetes was set with Normal distribution, mean = 250, SD = 30, with 97.5% of CI = 308.8 MYA, and crown age of Dothideomycetes with Normal distribution mean = 360, SD = 20, with 97.5% of CI = 399 MYA. The substitution models were selected based on jModeltest2.1.1; GTR+I+G for LSU, rpb2 and SSU, and TrN+I+G for tef1 (the model TrN is not available in BEAUti 1.10.2, thus we used TN93). Lognormal distribution of rates was used during the analyses with uncorrelated relaxed clock model. The Yule process tree prior was used to model the speciation of nodes in the topology with a randomly generated starting tree. The analyses were performed for 100 million generations, with sampling parameters every 10000 generations. The effective sample sizes were checked in Tracer v.1.6 and the acceptable values are higher than 200. The first 20% representing the burn-in phase were discarded and the remaining trees were combined in LogCombiner 1.10.2., summarized data and estimated in TreeAnnotator 1.10.2. Bars correspond to the 95% highest posterior density (HPD) intervals. The scale axis shows divergence times as millions of years ago (MYA).
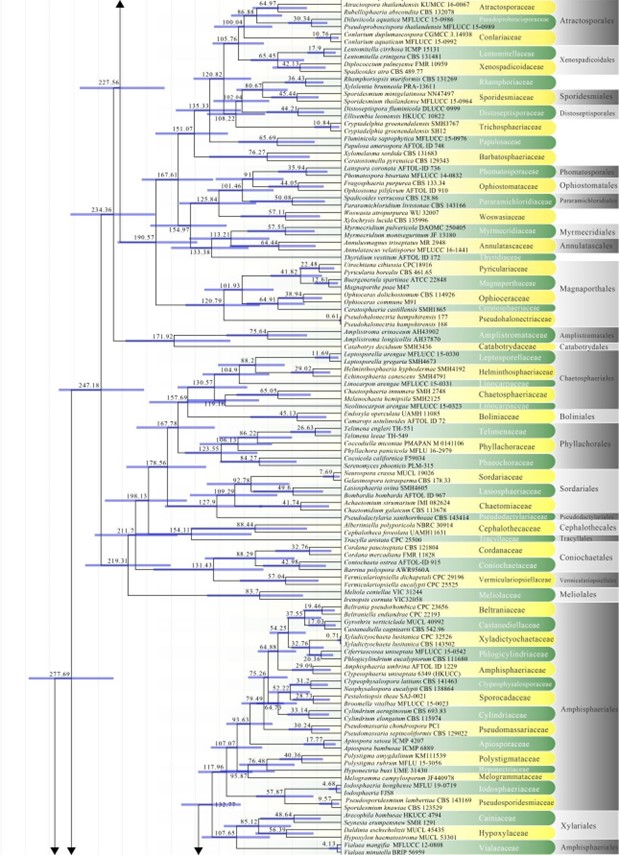
Figure 2 – Continued.
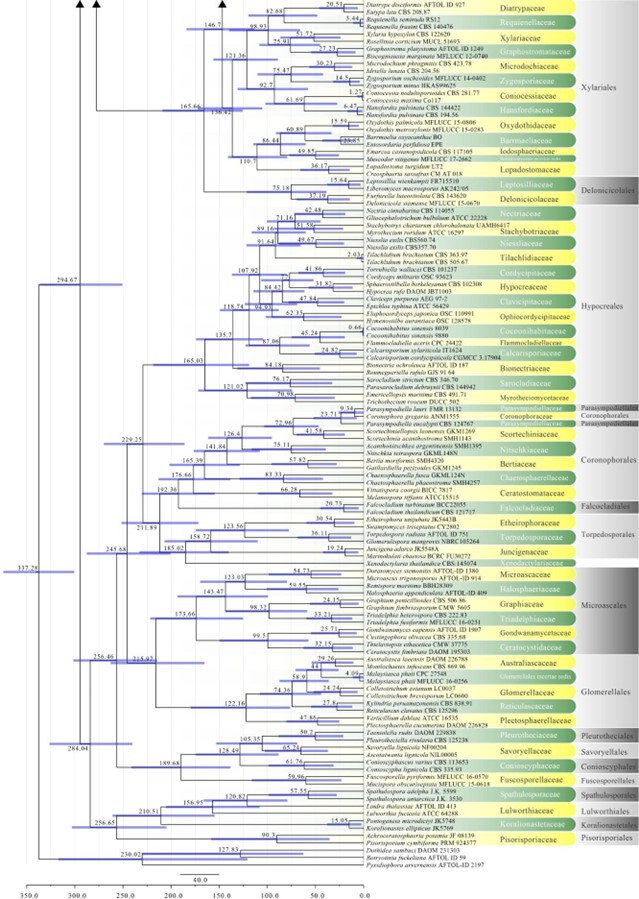
Figure 2 – Continued.
Orders
Sordariomycetidae families incertae sedi
Sordariomycetidae genera incertae sedis
