Spathulosporales Kohlm., Mycologia 65(3): 615 (1973)
Index Fungorum number: IF 90500; MycoBank number: MB 90500; Facesoffungi number: FoF 01798;
Spathulosporales includes Hispidicarpomycetaceae and Spathulosporaceae, while Hispidicarpomycetaceae lacks sequence data in GenBank. Sequence data are available only for two species of Spathulospora. Our phylogenetic analyses with combined LSU, SSU, and ITS data showed that Spathulospora and Rostrupiella species form a well-supported single clade (99% ML, 1.00 PP) within Lulworthiaceae (Fig. 16) and this is similar to Inderbitzin et al. (2004), Campbell et al. (2005) and Jones et al. (2009, 2019). However, sequence data are not available for the type species of Spathulospora hence, further collections are needed from the type to resolve the phylogenetic placement of this order. The divergence time for Sordariales is estimated at 121 MYA (Fig. 2), which falls within the range of family status. The order status may need revision following further study. Currently, there are two families and three genera in this order (this paper).
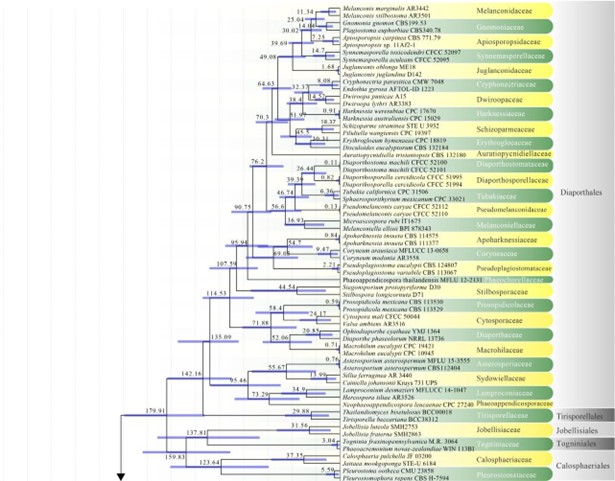
Figure 2 – The maximum clade credibility (MCC) tree, using the same dataset from Fig. 1. This analysis was performed in BEAST v1.10.2. The crown age of Sordariomycetes was set with Normal distribution, mean = 250, SD = 30, with 97.5% of CI = 308.8 MYA, and crown age of Dothideomycetes with Normal distribution mean = 360, SD = 20, with 97.5% of CI = 399 MYA. The substitution models were selected based on jModeltest2.1.1; GTR+I+G for LSU, rpb2 and SSU, and TrN+I+G for tef1 (the model TrN is not available in BEAUti 1.10.2, thus we used TN93). Lognormal distribution of rates was used during the analyses with uncorrelated relaxed clock model. The Yule process tree prior was used to model the speciation of nodes in the topology with a randomly generated starting tree. The analyses were performed for 100 million generations, with sampling parameters every 10000 generations. The effective sample sizes were checked in Tracer v.1.6 and the acceptable values are higher than 200. The first 20% representing the burn-in phase were discarded and the remaining trees were combined in LogCombiner 1.10.2., summarized data and estimated in TreeAnnotator 1.10.2. Bars correspond to the 95% highest posterior density (HPD) intervals. The scale axis shows divergence times as millions of years ago (MYA).
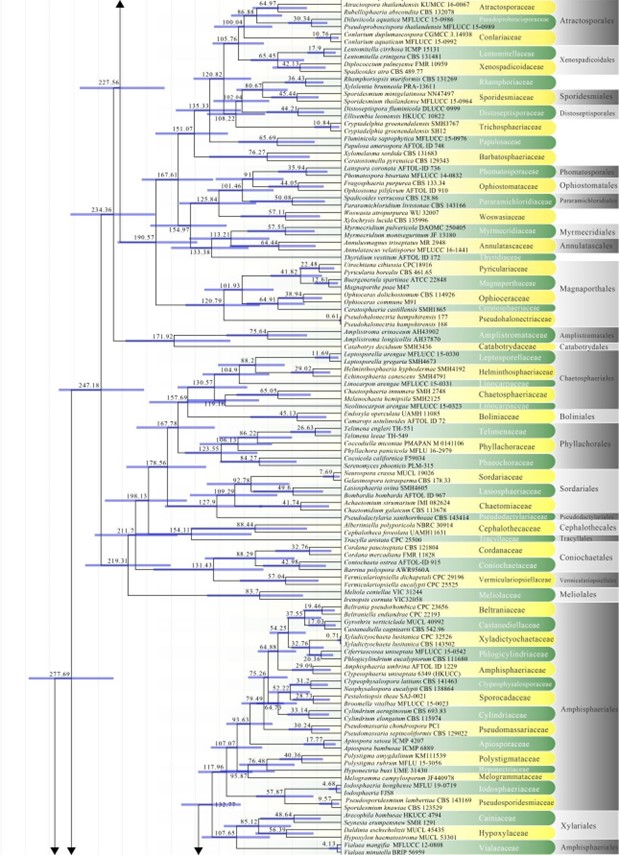
Figure 2 – Continued.
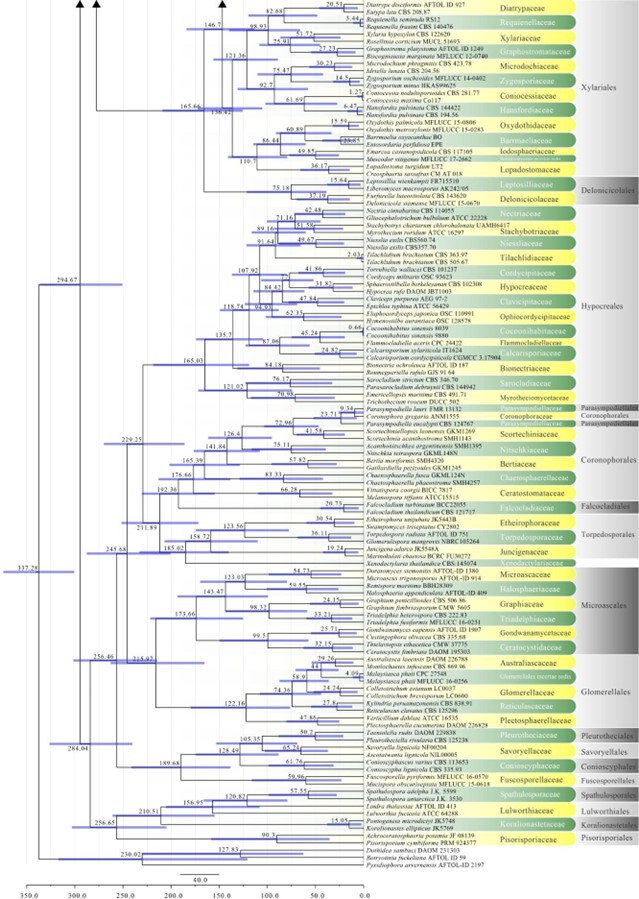
Figure 2 – Continued.
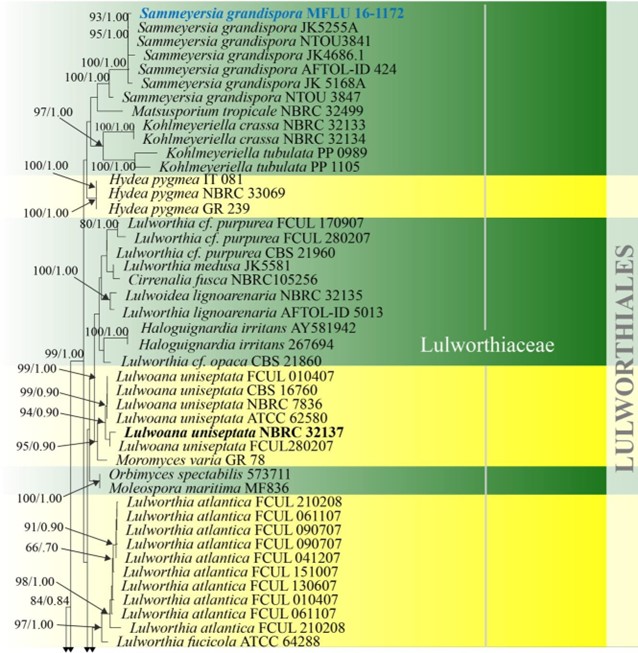
Figure 16 – Phylogram generated from maximum likelihood analysis based on combined LSU, SSU and ITS sequence data of selected taxa from Koralionastetales, Lulworthiales and Pisorisporiales. Sixty-nine strains are included in the combined gene analyses comprising 2780 characters after alignment (930 characters for LSU, 1059 characters for SSU and 784 characters for ITS). Fuscosporella pyriformis (MFLUCC 16-0570) and Parafuscosporella mucosa (MFLUCC 16- 0571) are used as outgroup taxa. Analyses of each single gene were performed and the topology of each tree had clade stability. The tree topology of the maximum likelihood was similar to the Bayesian and maximum parsimony analyses. Maximum likelihood analysis with 1000 bootstrap replicates yielded a best tree with the likelihood value of -24852.531353. The matrix had 1582 distinct alignment patterns, with 38.88% of undetermined characters or gaps. Estimated base frequencies were as follows; A = 0.249996, C = 0.239207, G = 0.291483, T = 0.219314; substitution rates AC = 1.016404, AG = 2.006035, AT = 1.090073, CG = 1.243791, CT = 5.331398, GT = 1.000000; gamma distribution shape parameter α = 0.395174. Maximum parsimony (black) and maximum likelihood (black) bootstrap values >65% and Bayesian posterior probabilities (green) >0.90 (ML/BYPP) are given above the nodes. Ex-type strains are in bold and new strains are in blue.
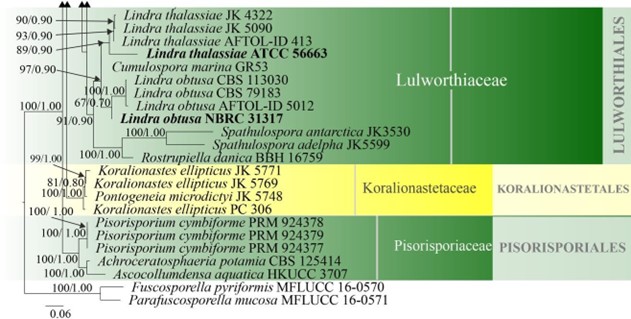

Figure 16 – Continued.
Families
