Vermiculariopsiellales Hern.-Restr., J. Mena, Gené & Crous, Stud. Mycol. 86: 91 (2017)
Index Fungorum number: IF 820346; MycoBank number: MB 820346; Facesoffungi number: FoF 12987;
Hernández-Restrepo et al. (2017) introduced Vermiculariopsiellales and one family, Vermiculariopsiellaceae, and one genus, Vermiculariopsiella, were accepted. ITS, LSU, SSU, actA and tef1 sequence data are available for species of Vermiculariopsiella. In our phylogenetic analyses generated from LSU, ITS, SSU and rpb2 sequence data, Vermiculariopsiellales formed a monophyletic clade with high statistical support (100% MPBS/1.00 PP) (Fig. 22). The divergence time for Vermiculariopsiellales is estimated at 131 MYA (Fig. 2). Currently, there is one family with one genus in this order (this paper).
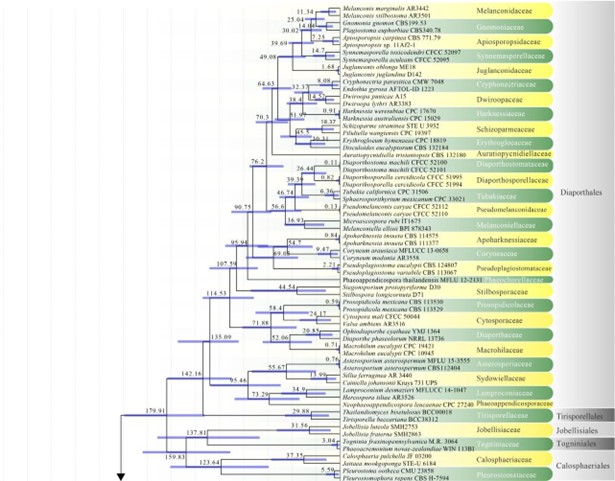
Figure 2 – The maximum clade credibility (MCC) tree, using the same dataset from Fig. 1. This analysis was performed in BEAST v1.10.2. The crown age of Sordariomycetes was set with Normal distribution, mean = 250, SD = 30, with 97.5% of CI = 308.8 MYA, and crown age of Dothideomycetes with Normal distribution mean = 360, SD = 20, with 97.5% of CI = 399 MYA. The substitution models were selected based on jModeltest2.1.1; GTR+I+G for LSU, rpb2 and SSU, and TrN+I+G for tef1 (the model TrN is not available in BEAUti 1.10.2, thus we used TN93). Lognormal distribution of rates was used during the analyses with uncorrelated relaxed clock model. The Yule process tree prior was used to model the speciation of nodes in the topology with a randomly generated starting tree. The analyses were performed for 100 million generations, with sampling parameters every 10000 generations. The effective sample sizes were checked in Tracer v.1.6 and the acceptable values are higher than 200. The first 20% representing the burn-in phase were discarded and the remaining trees were combined in LogCombiner 1.10.2., summarized data and estimated in TreeAnnotator 1.10.2. Bars correspond to the 95% highest posterior density (HPD) intervals. The scale axis shows divergence times as millions of years ago (MYA).
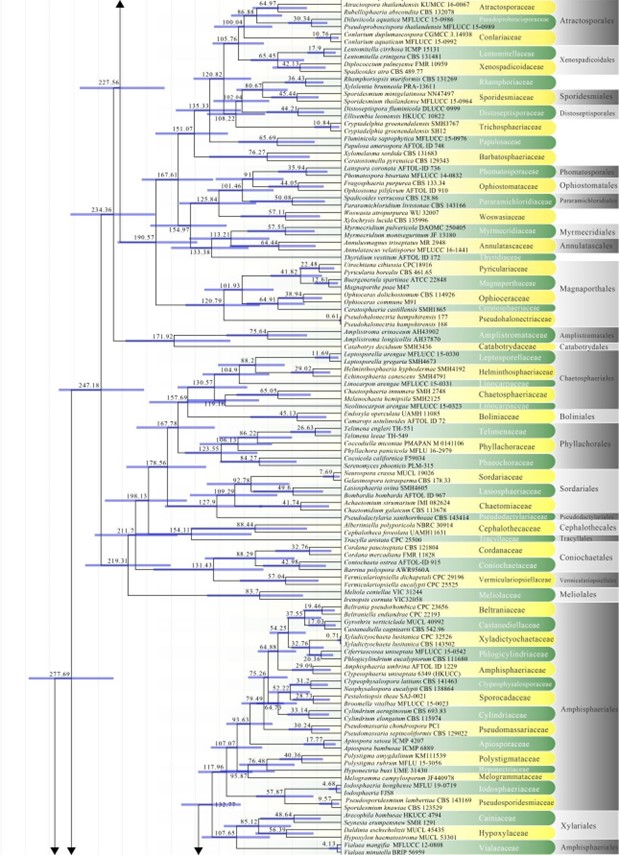
Figure 2 – Continued.
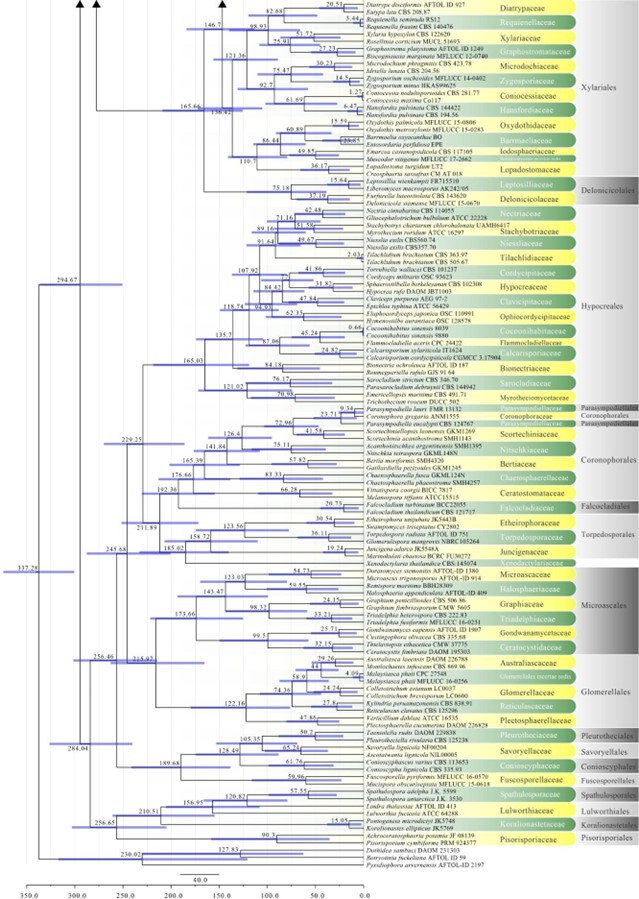
Figure 2 – Continued.
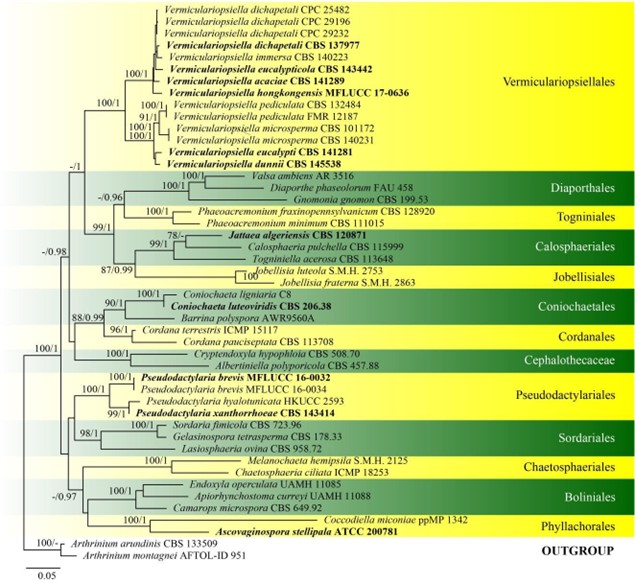
Figure 22 – Phylogram generated from maximum likelihood analysis based on combined LSU, SSU, ITS and rpb2 sequence data of Pseudodactylariales and Vermiculariopsiellales. Forty-seven strains are included in the combined analyses which comprised 4320 characters (865 characters for LSU, 1634 characters for SSU, 662 characters for ITS, 1159 characters for rpb2) after alignment. Arthrinium arundinis (CBS 133509) and Arthrinium montagnei (AFTOL-ID 951) (Apiosporaceae, Xylariales) are used as outgroup taxa. Single gene analyses were carried out and the phylogenies were similar in topology and clade stability. Tree topology of the maximum likelihood analysis is similar to the Bayesian analysis. The best RaxML tree with a final likelihood value of – 29256.470401 is presented. Estimated base frequencies were as follows: A = 0.249068, C = 0.239305, G = 0.280423, T = 0.231203; substitution rates AC = 1.369489, AG = 2.705642, AT = 1.345736, CG = 1.341396, CT = 6.599852, GT = 1.000000; gamma distribution shape parameter a = 0.220775. Bootstrap support values for ML greater than 75% and Bayesian posterior probabilities greater than 0.95 are given near the nodes, respectively. Ex-type strains are in bold.
Families
