Pseudocryptosphaerella S.K. Huang & K.D. Hyde, gen. nov.
Etymology: Name refers to genus being similar to Cryptosphaerella.
MycoBank number: MB 558359; Index Fungorum number: IF 558359; Facesoffungi number: FoF 10168;
Saprobic on wood. Sexual morph: Ascomata solitary or scattered, immersed to erumpent, black, turbinate, the apex collapsing when dry, tuberculate, sitting in a subiculum, lacking ostioles, with a central, conical Quellkörper. Peridium composed of brown to hyaline cells of textura angularis to textura prismatica, Munk pores present. Paraphyses absent. Asci 8- to multi-spored, unitunicate, clavate, with long pedicel, apex rounded, without apical ring, evanescent. Ascospores overlapping, hyaline, ellipsoidal cylindrical to broadly fusiform, slightly curved, aseptate, smooth- walled, guttulate. Asexual morph: Undetermined (adapted from Mugambi & Huhndorf 2010).
Notes – Mugambi & Huhndorf (2010) introduced and sequenced four ‘Cryptosphaerella’ species, C. costaricensis, C. cylindriformis, C. elliptica and C. malindensis. These species clustered in a clade and are characterized by lacking ostioles in the ascomata and a Quellkörper, polysporous asci and ellipsoidal to broadly fusiform ascospores (Mugambi & Huhndorf 2010). In this study, they cluster with 100%ML/1.00BY support (Fig. 1) and are sister to Biciliospora, Scortechiniella and Scortechiniellopsis in Scortechiniaceae. Therefore, we accept these species as members of Pseudocryptosphaerella based on phylogenetic result.
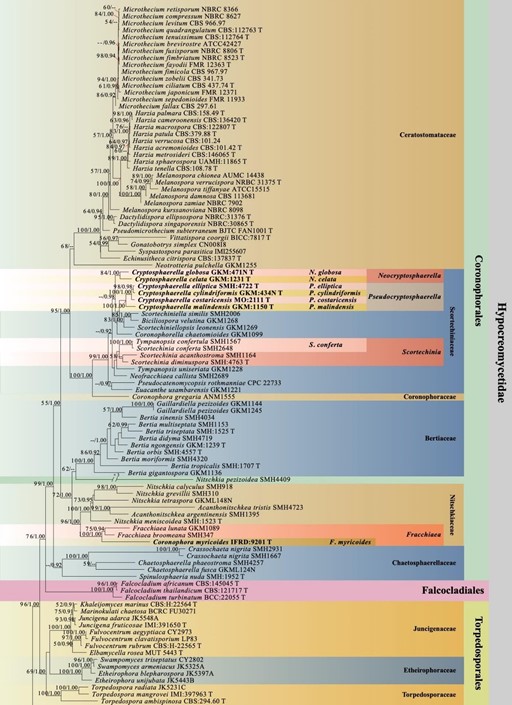
Figure 1 – Phylogram generated from maximum likelihood analysis based on combined LSU, SSU, TEF, RPB2, TUB and ITS sequence data with the confidence values of bootstrap (BS) proportions from the Maximum Likelihood (ML) analysis (ML-BS > 50%, before the backslash) and the posterior probabilities (PP) from the Bayesian (BY) analysis (BY-PP > 0.90, after the backslash) above corresponding nodes. The ‘–’ indicates a lack of statistical support (< 50% for ML-BS and < 0.90 for BY-PP). One hundred and ninety-eight strains are included in the combined analyses which comprised 4581 characters (842 characters for LSU, 1025 characters for SSU, 950 characters for TEF, 876 characters for RPB2, 391 characters for TUB, 497 characters for ITS) after alignment. Strains of Xylariomycetidae are used as outgroup taxa. The best score in IQ-TREE explores with a final likelihood value of -90984.2293 is presented. The model of each partitioned gene is LSU: GTR+I+G; SSU: GTR+G; TEF: GTR+I+G; RPB2: GTR+I+G; TUB: GTR+I+G; ITS: GTR+G. Sequences generated indicated in bold. Strain numbers are noted after the species names and ex-type strains marked with ‘T’ after the culture number. Alignment is available at TreeBASE (URL: http://purl.org/phylo/treebase/phylows/study/TB2:S28271).
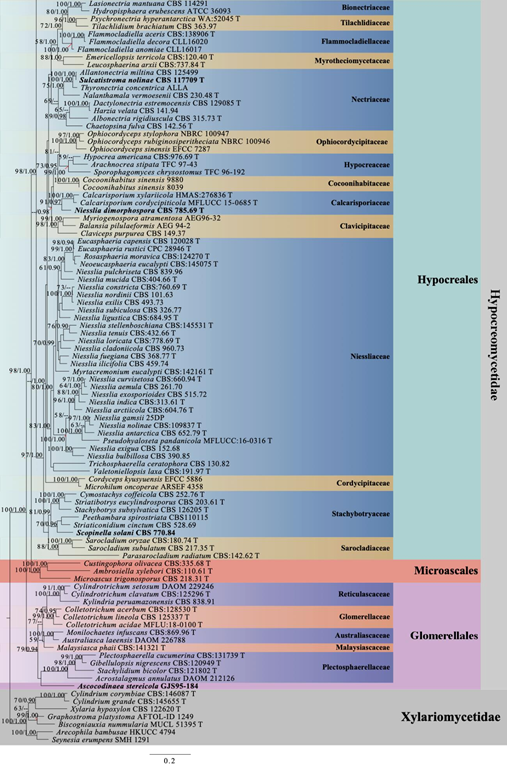

Figure 1 – Continued.
Species
