Plectosphaerella cucumerina (Lindf.) W. Gams, in Domsch & Gams, Fungi in Agricultural Soils: 160 (1972)
MycoBank number: MB 320609; Index Fungorum number: IF 320609; Facesoffungi number: FoF 12962;
Associated with leaf spots of Bridelia sp. Sexual morph: Vegetative hyphae 1.4–3.0 µm wide (x̄ = 1.88, n = 40), hyaline to brown, smooth-walled, septate, branched. Ascomata developing on MEA, 145–150 × 120–147 μm (x̄ = 148×130 μm, n = 5), superficial or immersed in culture colonies, scattered, solitary to gregarious, subglobose to ovoid, uni- or multi-loculate. Peridium 18–32 μm (x̄ = 24.2 μm, n = 10) wide, composed of 1–3 layers, inner layers comprising hyaline to dark brown, pseudoparenchymatous cells of textura angularis, outer layers composed of thick, dark brown to black, cells of textura angularis. Asci 42–58 × 6–13 μm (x̄ = 50.7 × 8.92 μm, n = 10), 8-spored, unitunicate, cylindrical to obpyriform, thick-walled at apex, with J-, sub-apical ring. Ascospores 6.75–14.0 × 2.00–3.65 μm (x̄ = 7.46 × 2.85 μm, n = 30), hyaline, cylindrical to clavate or fusiform, with rounded apices and bases, smooth-walled, 1-septate, guttulate, slightly constricted in the middle. Asexual morph: Conidiophores reduced to conidiogenous cells. Conidiogenous cells 6.50–10.5 × 4.80–6.00 μm (x̄ = 8.20 × 5.57 μm, n = 5), enteroblastic, phialidic, solitary, ellipsoidal to ovoid, smooth-walled, hyaline to pale brown. Appressoria not observed.
Culture characteristics: Colonies on MEA reaching approximately 70 mm diam. after 5 days at 25 °C, flat, entire margin, aerial mycelia, dense, fast-growing, pale orange with hyaline mycelium, reverse yellowish white. Sporulate after 20 days.
Material examined: THAILAND, Chiang Rai Province, Omkoi, on living leaves of Bridelia sp. (Phyllanthaceae), 16 October 2019, Gomdola D., (MFLU XXX), living culture MFLUCC XXX.
Hosts and distribution: Foeniculum vulgare, Helianthus annuus, Cucumis sativus, Medicago sativa, Lagenaria siceraria, Brassica oleracea, Lycopersicon esculentum, Phaseolus vulgaris, Platostoma palustre, Sedum sp., Solanum lycopersicum from China; Arabidopsis thaliana, Pyrus malus from Switzerland; Bambusa sp. from Iran; Dieffenbachia maculate, Phlox sp. from New Zealand; Galium spurium, Solanum melongena, Pisum sativum from Canada; Nicotiana tabacum from Belgium; Solanum tuberosum from Pakistan; Viola odorata from Egypt; Viola tricolor from Netherlands; Asparagus officinalis, Aquilegia flabellata, Diplotaxis tenuifolia, Capsicum annuum, Cichorium endivia, Citrullus lanatus, Cucumis melo, Ocimum basilicum, Valerianella locusta, Petroselinum crispum, Solanum lycopersicum from Italy; Glycine max, Heterodera glycines, Heuchera sanguinea, Ocimum basilicum from USA (Farr and Rossman 2022).
Notes: Our collection clustered with the strains of Plectosphaerella cucumerina (CBS 133.37, CBS 133.33, CBS 101014, CBS 632.94, CBS 139.60, CBS 355.36, CBS 619.74, CBS 367.73, CBS 286.64, CBS 400.58, CBS 101.958, CBS 567.78) in the combined ITS, LSU, β-tubulin and tef1-α locus analysis with strong bootstrap support value (ML/MP/BI = 98%/98%/1.00). Morphologically, our collection fits to Plectosphaerella cucumerina by its sexual morph (Carlucci et al. 2012). Therefore, we identified our collection as Plectosphaerella cucumerina based on morphological and molecular evidences.
There are several reports about disease profile of P. cucumerina that cause necrotic leaf spots on Aquilegia flabellata in Italy (Garibaldi et al. 2021), Alfalfa root rot in China (Zhao et al. 2021), Fennel root rot in China (Cai et al. 2021), and Chinese cabbage wilt in China (Gao et al. 2022). Here, our P. cucumerina collection associated with leaf spots of Bridelia sp. This is the first report of Plectosphaerella cucumerina on living leaves of Bridelia sp. from Thailand and it is a new host and geographical record.
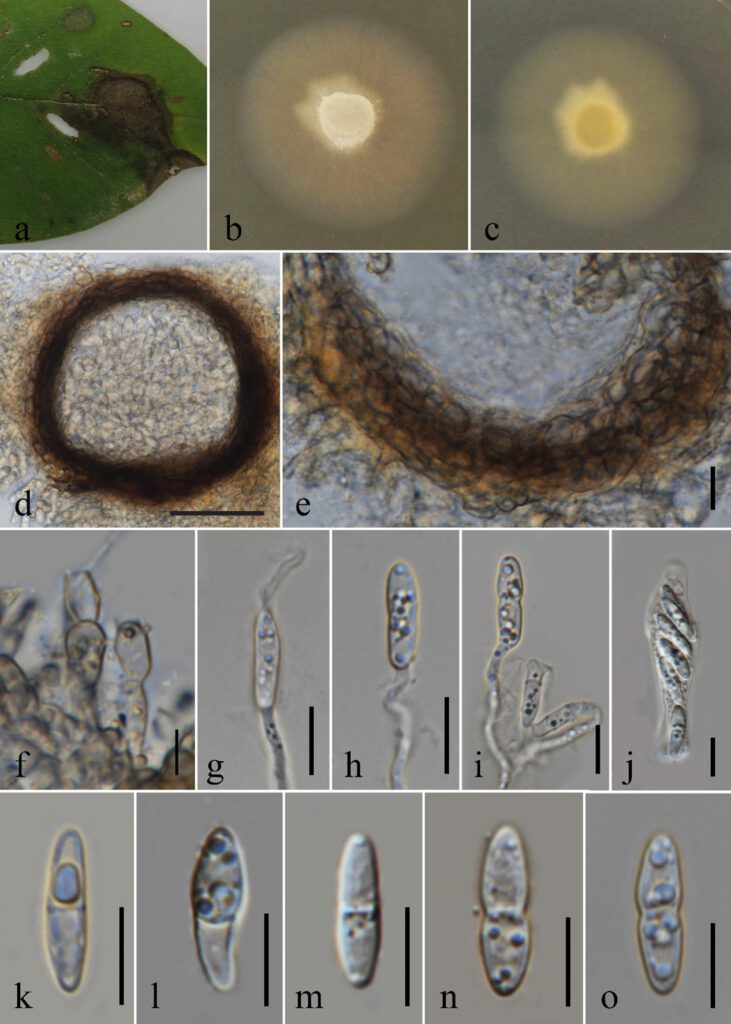
Fig. xx. Plectosphaerella cucumerina a Examined materials with leaf spots. b Surface view of colony on MEA. c Reverse view of colony on MEA. d Section through ascomata. e Peridium. f Conidiogenesis g–i Germinating ascospores. tubes emerging from conidium j Asci. k–o Ascospores. Scale bar. d = 50 μm e, g-o = 10 μm f = 5 μm.
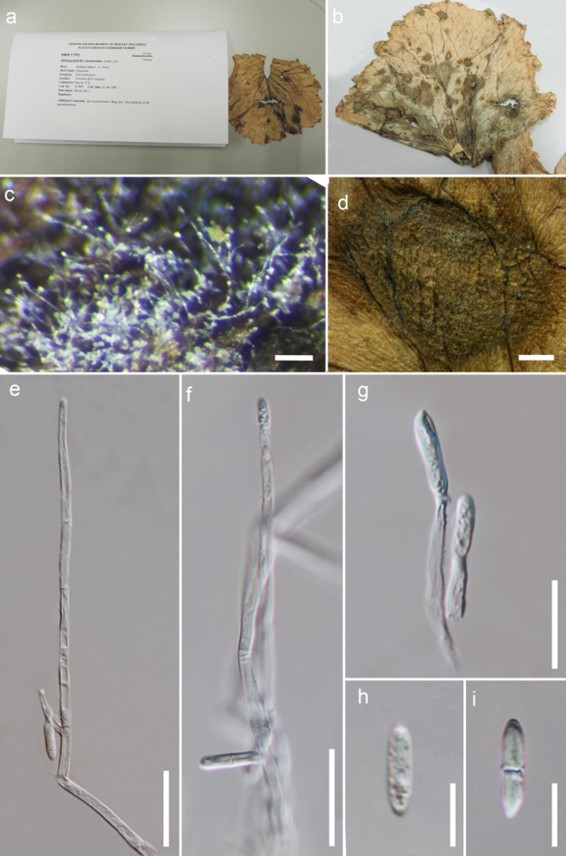
Figure 201 – Plectosphaerella cucumerina (Material examined – AUSTRALIA, Queensland, Tewantin, on the leaf of Lactuca sativa, 22 January 1991, Mayers, P.E., BRIP 17392, = Monographella cucumerina). a Herbarium material. b Host. c Colony on the host. d Appearance of colonies on leaf surface. e, f Conidiophores. g Conidiogenous cell. h, i Conidia. Scale bars: c = 100 µm, d = 1 mm, e, f = 25 µm, g = 15 µm, h-i = 5 µm.
