Cryphonectria kunmingensis Lu L., K.D. Hyde & Tibpromma S., sp. nov. Fig. xx
MycoBank number: MB; Index Fungorum number: IF; Facesoffungi number: FoF 10843;
Etymology: based on the location where the fungus was collected.
Saprobic on dead stem of unknown plant. Sexual morph Ascostromata (excluding neck) 300–660 × 240–420 μm (x̅ = 526 μm, n = 342), single, semi-immersed, orange, oval to irregular. Perithecia immersed, single or aggregated, subglobose or irregular, multi-loculate, with long necks, ostiolar canal sometimes immersed in ascostromatic tissues, or come out from substrate, orange. Neck composed of umber stromatic cells of textura porrecta. Peridium consists of two distinct layers, cells of textura angularis, outer layer dense, thick, yellow to pale brown, inner layer composed of 4-5layers, hyaline. Hamathecium 6–9 μm wide (x̅ = 7 μm, n = 20) consists of a cellular, septate, hyaline, paraphyses attached at base. Asci 55–70 × 8–10 μm (x̅ = 64.5 × 8.9 μm, n = 20), 8-spored, unitunicate, cylindrical, hyaline, with distinct J- refractive ring, no stalked. Ascospores 9.5–15 × 3–5 μm (x̅ = 13 × 4.2 μm, n = 20), overlapping uniseriate to biseriate, ellipsoid, 1–2-septate when maturity, constricted at septa, hyaline, smooth-walled, without mucilaginous sheath. Asexual morph Undetermined.
Culture characteristics: colonies on PDA reaching 60 mm diam after 4 weeks in 20 °C, circular, flat, lobate, pale yellow, flocculent, a lot of aerial mycelia front view pale yellow, reverse view orange center turning pale yellow towards the margin, produced pigments in culture.
Material examined: CHINA, Yunnan Province, Kunming, Chang chong Mountain, on dead stem of unknown plant, 20 June 2021. Li Lu, CCS 4, (HKAS 121976, holotype), ex-type culture, KUMCC 21–0216.
GenBank accession numbers: ITS = xxxxxx, LSU = xxxxxxx, TUB1 = xxxxxxx, rpb2 = xxxxxxx, tef1-α = xxxxxxx
Notes: In the combined ITS, LSU, rpb2, TUB1 and tef1-α gene analysis showed that our collection formed a distinct subclade (Fig. xx) with high bootstrap support (MP/BI=98/1.00). ITS sequence of our strain is 93.6% similar to Cryphonectria nitschkei (GQ290656) while LSU is 99.6% similar to C. nitschkei (AF408341). The blast results of tef1-α is 90% identical to C. macrospora (AY308952), rpb2 is 96.5% identical to Chromendothia citrina (DQ862015) and TUB1 is 92% identical to Cryphonectriaceae sp. (JQ862910). Morphologically, our collection is similar to general characters of Cryphonectria (Jiang et al. 2020), however, it differs from other Cryphonectria species in having 2-septated ascospores, in addition, our ascospores (9.5–15) are longer compared to the morphologically similar species C. nitschkei (8.5–12.5) (Myburg et al. 2004). Therefore, we introduce our collection as a new species, Cryphonectria kunmingensis based on morphology and phylogeny. There are four species have been collected from China viz. C. japonica, C. neoparasitica, C. quercus and C. quercicola and this is the fifth species collected in China.
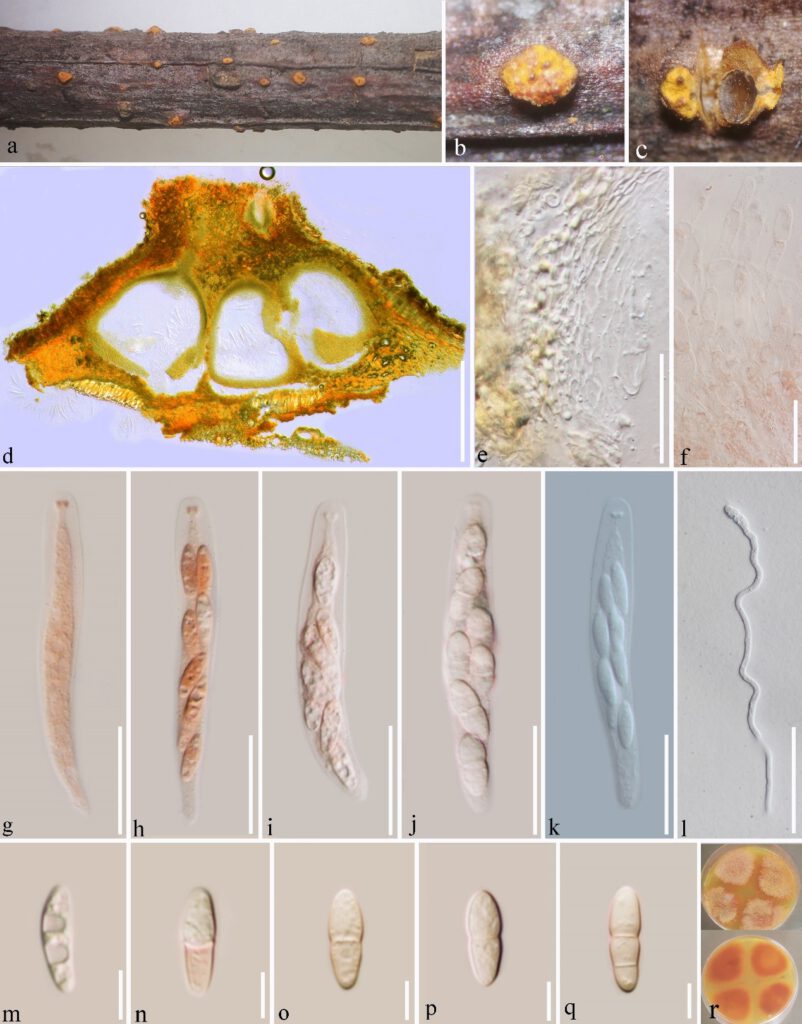
Fig. 1. Cryphonectria kunmingensis (HKAS 121976, holotype). a-c Ascostromata on the host surface. d Cross section of ascostromata. e Peridium. f Paraphyses. g-j Asci (stained in Congo Red reagent). k J- refractive ring (stained in Melzer’s reagent). l Germinated ascospore. m-q Ascospores (stained in Congo Red reagent). r Upper and reverse views of colonies on PDA. scale bars: d = 300 µm, e = 30 µm, f = 25 µm, g-l = 20 µm, m-q = 5 µm.
