Translucidithyrium chinense H.X. Wu & X.H. Li, in Li et al., MycoKeys 76, 7 (2020)
Index Fungorum number: IF 557843; Facesoffungi number: FoF 09429; Fig. 4
Epiphyllous appearing as brownish lesions on the surface of unidentified leaves. Mycelium scarce or not distinct. Sexual morph: Ascomata 600–700 μm in diam. (x̅ = 650 μm, n = 15), solitary, scattered, globose to subglobose, flattened, light brown, without ostiole. Peridium poorly developed at the basal, thin at the upper walls, membranous, composed of pale brown, interwoven, irregular, meandering, interwoven arranged cells. Hamathecium composed of hyaline interascal tissues. Asci 55–70 × 40–70 μm (x̅ = 65 × 50 μm, n = 15), bitunicate, 8-spored, globose to subglobose, embedded in interascal tissues, apedicellate. Ascospores 35–40 × 10–15 μm (x̅ = 37 × 12 μm, n = 20), crowded, irregularly overlapping, hyaline, ovoid at young state, fusiform to inequilateral at maturity, tapering at both ends, 1-septate, constricted at the septum, lower cell longer than the upper, hook-like curve in the upper cell, verrucose to smooth, covering a thin sheath. Asexual morph: Undetermined.
Material examined – Thailand, Nan Province, Pua District, unidentified leaf, 4 February 2022, Diana Sandamali, MFLU 23-0065.
GenBank number – ITS: OQ568947, LSU: OQ568948.
Known distribution (based on molecular data) – China (Li et al. 2020), Thailand (this study).
Known hosts (based on molecular data) – Alpinia blepharocalyx (Li et al. 2020).
Notes – The morphology of this species resembles Translucidithyrium chinense in having brownish, globose ascomata, globose to subglobose, 8-spored asci, and hyaline, 1-septate, curved ascospores that are initially ovoid, becoming fusiform at maturity (Li et al. 2020). The comparisons of the ITS sequences show four base pair differences across 450 bp (1.1 %). Our ITS and LSU combined phylogenetic analyses (Fig. 3) reveal that our strain clusters with Translucidithyrium chinense (IFRDCC 3000) with 100% maximum likelihood and 1.00 posterior probability support. Thus, we report the first geographical record of T. chinense in Thailand.
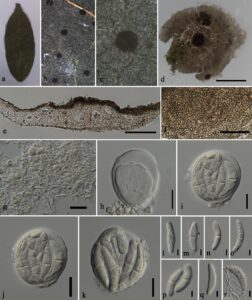
Fig. 1 – Translucidithyrium chinense (MFLU 23-0065, new geographical record). a Leaf specimen. b, c Ascomata on the leaf surface. d Squash mount of ascomata. e Section through ascomata. f Interascal tissue. g Upper cell wall. h Immature ascus. i–k Mature asci. l–q Immature to mature ascospores. r Germinated ascospore. Scale bars: c = 200 µm, d = 100 μm, e, g–j = 20 μm, f, k–r = 10 μm.

Fig. 2 – Phylogram generated from maximum likelihood analysis based on combined LSU and ITS sequence data. Thirty-nine strains are included in the combined gene analyses comprising 1462 characters after alignment (893 characters for LSU and 569 for ITS). Dothidea sambuci (AFTOL ID-274) is used as the outgroup taxon. The tree topology of the Bayesian analysis was similar to the maximum likelihood analysis. The best RaxML tree with a final likelihood value of -12266.208 is presented. The matrix had 768 distinct alignment patterns, with 19.62% undetermined characters or gaps. Evolutionary models Tn+F+R6 and GTR+G were selected as the best-fit models for the LSU and ITS gene regions, respectively. Bootstrap values for maximum likelihood equal to or greater than 60 and Bayesian posterior probabilities equal to or greater than 0.90 are placed above or below the branches, respectively. Ex-type strains are in bold. The newly generated sequences are indicated in yellow.
