Kirschsteiniothelia extensum. S. Wang, Q. Zhao & K.D. Hyde sp. nov. Fig. 9
Mycobank number: MB; Index Fungorum number: IF; Facesoffungi number: FoF 11803;
Etymology:
Holotype: X
Saprobic on decaying wood. Sexual morph: Not observed. Asexual morph: Colonies effuse, scattered, brown or black, hairy, glistening. Mycelium partly superficial, partly immersed in the substratum, composed of brown, septate, branched hyphae. Conidiophores macronematous, mononematous, solitary, cylindrical, straight or slightly flexuous, dark brown, unbranched, thick-walled,smooth, slightly tapering towards the apex, 4–9 septate, truncate at the apex, 80–230 µm (x̄ = 140 µm, n = 15) long, 6.5–9.5 μm (x̄ =7.5 µm, n = 15) wide. Conidiogenous cells integrated, terminal, monoblastic, percurrent, pale brown, cylindrical, 11–19 µm (x̄ = 15 µm, n = 15) long, 4–7.5 μm (x̄ =6 µm, n = 15) wide. Conidia acrogenous, solitary, smooth, obclavate, straight or slightly curved, tapering to the apex, 5–8-euseptate, becoming pale brown to pale towards the apex, truncate at the base, 45–120 µm (x̄ = 60 µm, n = 30) long, 5–12 μm (x̄ = 9 µm, n = 30) wide.
Culture characteristics: Conidia germinating on MEA within 12h. at room temperature. Colonies on MEA, unregular round shape, becoming filamentous in the culture, mycelium slightly raised, flattened, filiform, gray aerial hyphae, spreading from the center, becoming olivaceous-brown at the surface and olivaceous-brown to black in reverse, MEA change to light gray to olivaceous-brown.
Material examined: Thailand, Nang Lae, Mueang Chiang Rai, Chiang Rai Province, saprobic on decaying wood, July 2020, Rongju Xu, MD73, (21-0130, holotype), extype culture, *****.
Notes: Kirchsteiniothelia extensum is introduced here based on both morphology and molecular data. Kirchsteiniothelia extensum forms a distinct clade within Kirschsteiniotheliaceae and is sister to K. submersa. The difference between them is conidiophores of Kirschsteiniothelia extensum (80–230 × 6.5–9.5 μm) are much shorter than those of K. submersa (220–280 × 6–7μm).
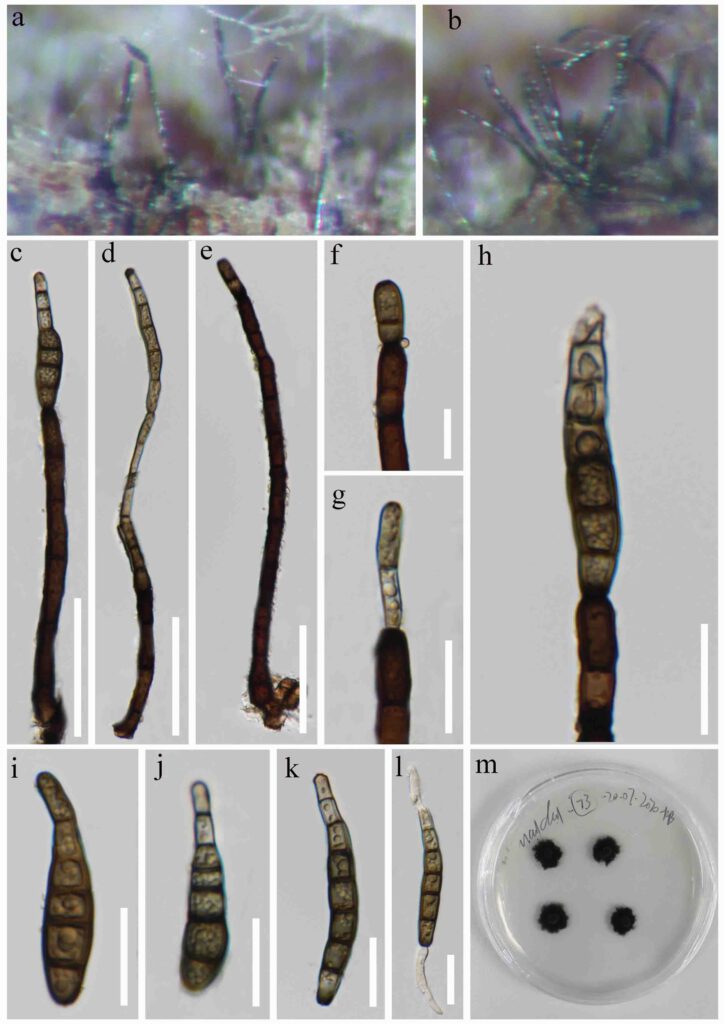
Fig. *** Kirschsteiniothelia extensum (MFLU ****, holotype) ab Colonies on dead wood. cd Conidiophore with conidia. e Conidiophore. f–h Conidiogenous cells and conidia. i–k Conidia. l Germinating conidium. m Culture on MEA Scale bars: c –e = 50 μm, f–l = 20 μm.
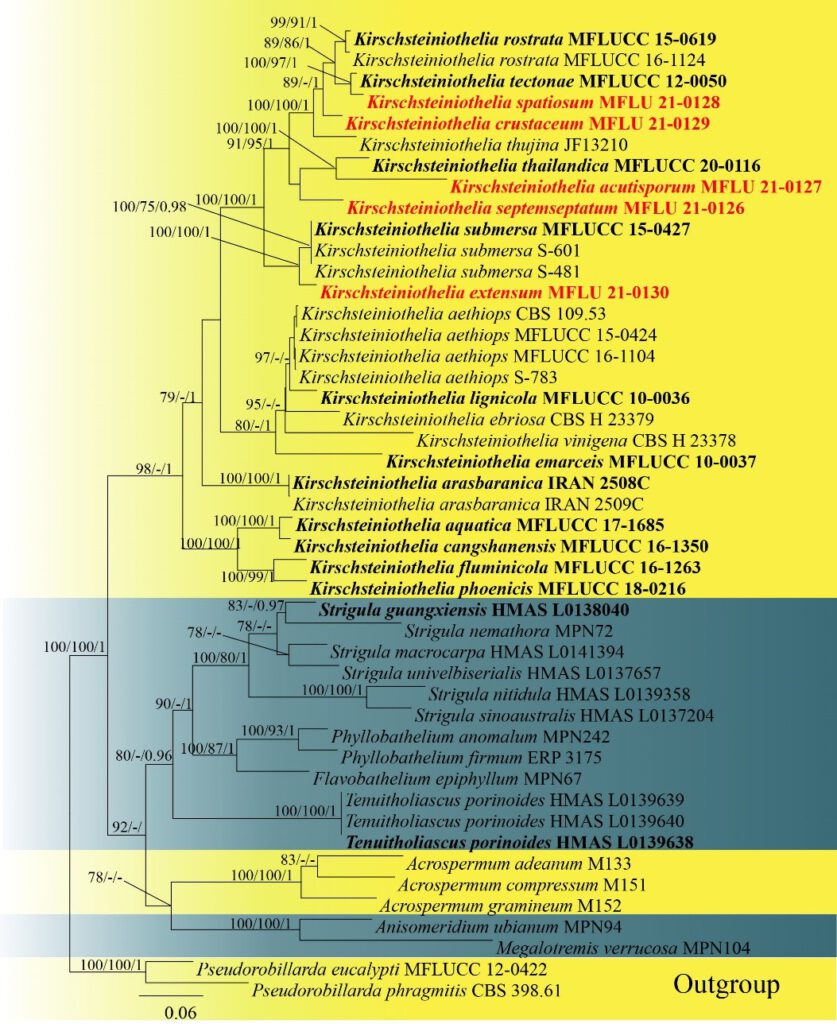
Figure 2. Phylogram generated from maximum likelihood analysis based on combined LSU, SSU,ITSsequence data. Forty-six taxa were included in the combined analyses, which comprised 2,104 characters (LSU = 1-788 bp, SSU= 789-1,632 bp, ITS = 1,633-2,104 bp), including alignment gaps. Among them, 1,191 characters were constant, 239 characterswere singleton sites,and 674characters were parsimony informative. The best scoring RA × ML tree with a final likelihood value of -16,278.480 is presented. Bootstrap support values for ML and MP equal to or greater than 75% and BYPP equal to or greater than 0.95 are given above the nodes. Pseudorobillarda eucalypti (MFLUCC 12–0422) and P. phragmitis(CBS 398.61)were used as the outgroup taxa. The newly generated sequence is indicated in red.The ex-type strains are indicated in bold.
