Dyfrolomyces distoseptatus M. Niranjan and V.V Sarma,sp. nov.
MycoBank number: MB 556726; Index Fungorum number: IF 556726; Facesoffungi number: FoF 06625; Fig. 30
Etymology: In reference to the distoseptate ascospores.
Saprobic on undetermined decaying twig. Sexual morph: Ascomata 550–630 high × 450–600 μm wide, perithecial, immersed in periderm, erumpent neck with pseudoparaphyses, clypeate, ostiolate, papillate. Peridium 40 μm, with two strata, outer thick, carbonaceous and inner brown and hyaline cells of textura angularis to epidermoidea cells. Hamathecium comprising 1.8–2.1 μm wide, filamentous, septate, unbranched, dense, narrow cellular pseudoparaphyses, longer than asci. Asci (123.1)126.7–146.2 (148.6) × (4.1)4.7–6.3 (6.5) μm (x̄ = 136.8 × 5.6, n = 25), 8–spored, bitunicate, cylindrical, apical ends obtuse, smooth-walled, short pedicellate, persistent. Ascospores (19.4)19.7–24.9 × (4.1) 4.3–5 μm ( x̄ = 21.9 × 4.7, n = 25), uni-seriate, fusoid, obtuse ends, hyaline, 3-distoseptate, apical ends slightly bent. Asexual morph: Undetermined.
Material examined: India, Andaman and Nicobar Islands, South Andaman, Manjery, (11˚52’25.7”N 92˚64’89.9”E), on unidentified twig, 10 December 2017, M. Niranjan (AMH 9984, holotype), extype living culture NFCCI: 4377.
GenBank numbers: ITS: MK024391, LSU: MH971236.
Notes: Dyfrolomyces distoseptatus clustered with D. thamplaensis with strong bootstrap support (100%) in ML analysis. It also forms a sister relationship clade with D. sinensis (Fig. 29). However, D. distoseptatus has several distinguishable characters. It is closely related to D. thamplaensis in having distoseptate ascospores with acute ends, but the ascospores in the latter have prominent guttules. In addition, the asci of D. distoseptatus are smaller than D. thamplaensis (123.1–148.6 × 4.1–6.3 μm vs. 114–160 × 6–8.5 μm) (Zhang et al. 2017a). Furthermore, D. distoseptatus produces ascospores that are 3-septate when compared to 6–7 septate ascospores in D. sinensis (Hyde et al. 2018). Hence, based on the above-mentioned morphological differences (Fig. 30) and molecular sequence analyses (Fig. 29), a new species, Dyfrolomyces distoseptatus, is introduced.
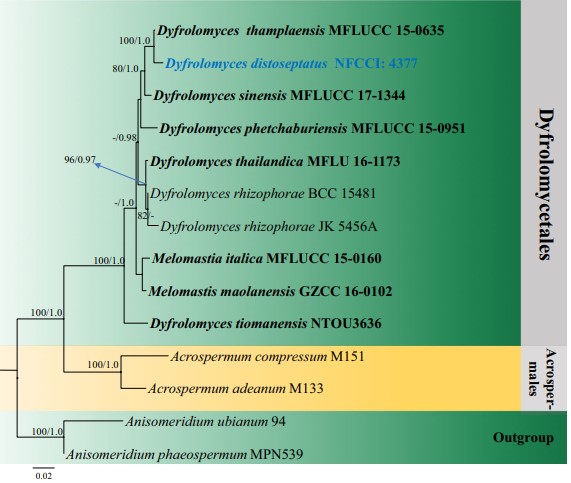
Fig. 29 Phylogram generated from maximum likelihood analysis (RAxML) of Dyfrolomycetales based on LSU and SSU sequence data. Maximum likelihood bootstrap values equal or above 70%, Bayesian posterior probabilities equal or above 0.90 (MLBS/PP) are given at the nodes. An original isolate number is noted after the species name. The tree is rooted to Anisomeridium phaeospermum (MPN539) and A. ubianum (94). The ex-type and reference strains are indicated in bold. Hyphen (-) represents support values below 70% MLBS and 0.90 PP
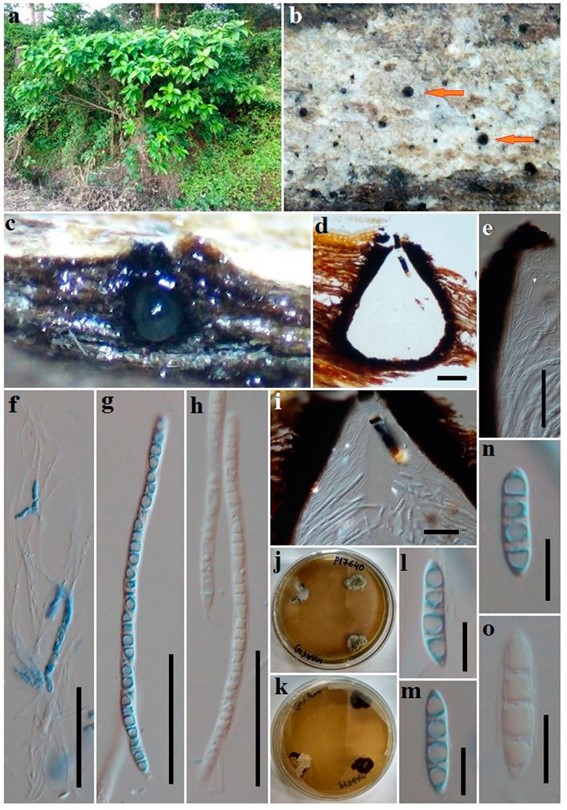
Fig. 30 Dyfrolomyces distoseptatus (AMH 9984, holotype). a Host plant. b Papillate necks. c, d, i Vertical section of ascomata. e Peridium. f Pseudoparaphyses. g, h Asci j, k Cultures in Petri plates from single spore isolations. l–o Ascospores. Scale bars: d = 100 µm, e–I = 50 µm, l–o = 10 µm
Species
