Colletotrichum plurivorum Damm, Alizadeh & Toy. Sato, in Damm, Sato, Alizadeh, Groenewald & Crous, Stud. Mycol. 92: 31 (2019)
MycoBank number: MB 824228; Index Fungorum number: IF 824228; Facesoffungi number: FoF 10691;
Endophytic were isolated from C. grandis cv ‘Tomentosa’ leaf. Sexual morph on PDA. Asci 30–70 × 10–20 μm, n= 50, lunt circle, hyaline, eight ascospore that were clavate. Ascospores 15–20 × 4–6 μm (x̄= 17 × 5, n= 50), hyaline, smooth-walled, aseptate, slightly curved, cylindrical with slightly sharp ends. Asexual morph on PDA. Pycnidium 60–160 × 60–200 μm (x̄ = 90 × 90, n= 30), spherical, chestnut, endophytic hyaline conidia. Conidia 10–20 × 5–7 μm (x̄= 16 × 6, n= 50), ellipsoid to stick, hyaline, smooth-walled, aseptate, the apex and base rounded. Appressoria was solitary or in loose groups, grey ollvaceous, oblong to serrated, edge integrity.
Cultural characteristics. Colonies on PDA reach 80 mm diam. in seven days at 28℃, growth rate 10–11 mm/day (x̄= 11 mm, n=6). Colonies circular, margin entire, flat, at first white and increasingly becoming dark mouse grey pigment deposition and elevation flat, colony from above: dark grey, reverse white to dull green to pale green. The culture emerged fruiting body and fulvous conidiomata at 4–6 weeks.
Material examined. China, Guangdong province, Huazhou, isolated from healthy leaf of C. grandis cv ‘Tomentosa’, May 2019, YX Shu, (dried culture ZHKU 21-0087), living culture ZHKUCC 21-0102).
Habit and host. Abelmoschus esculentus, Arundina graminifolia, Amorphophallus rivieri, Camellia sinensis, Carica papaya, Capsicum sp., Coffea sp., Cymbidium hookerianum, Gossypium sp., Mangifera indica, Manihot esculenta, Musa sp., Oncidium sp., Passiflora edulis, Phaseolus lunatus, Phaseolus vulgaris, Pyrus bretschneideri, Solanum lycopersicum, Soybean, Spathiphyllum wallisii.
Known distribution. Benin, Brazil, China, India, Iran, Japan, Mexico, Myanmar, Mylaysia, Taiwan, Thailand, Viet Nam.
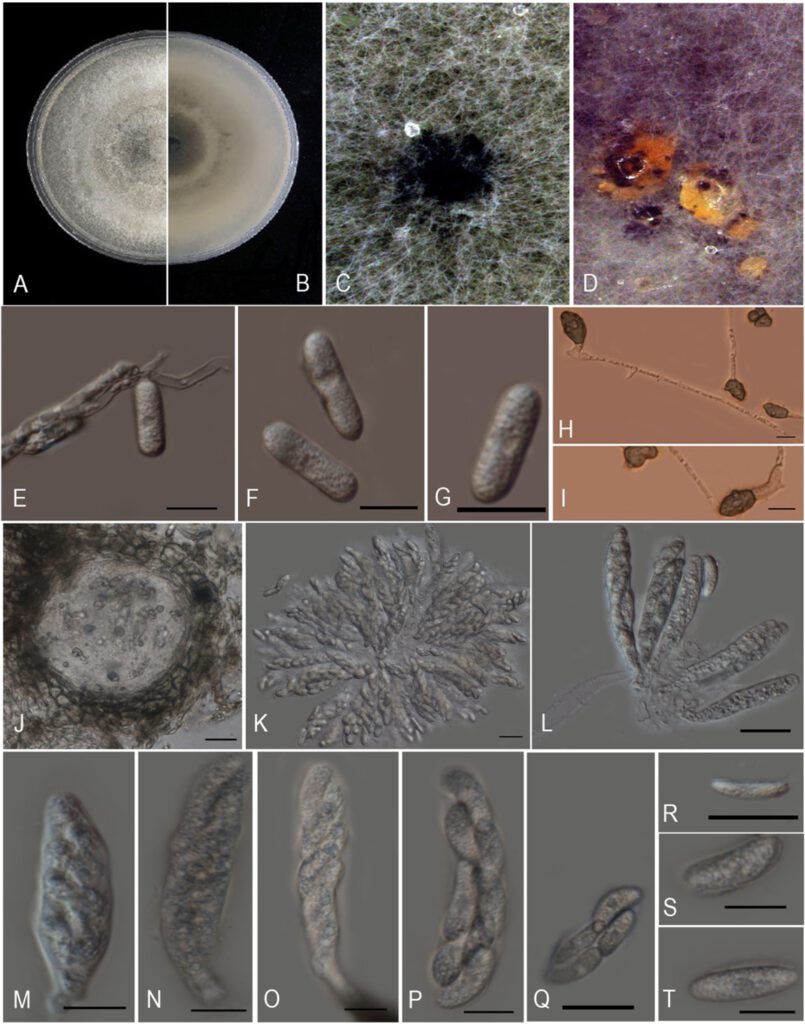
Morphological characteristics of Colletotrichum plurivorum (ZHKUCC 21-0102). A, B. Upper and reverse sides of cultures on PDA 7days after inoculation. C. fruit body. D. Conidiomata. E–G. Conidia on PDA. H–I. Appressoria. J. Pycnidium. K–P. Ascus. Q–T. Ascospore. Scale bars: 10 μm (E–G, M–P, S–T); 20 μm (H–K, Q–R).
