Synnemellisia punensis K. S. Pawar, P. N. Singh, S. K. Singh sp. nov.
MycoBank number: MB 843687; Index Fungorum number: IF 843687; Facesoffungi number: FoF 10810; Figs.1–3
Etymology: specific epithet ‘punensis’ refers to name Pune (a district of Maharashtra state).
Holotype: NFCCI 5166
Colour code follows: Methuen Handbook of Colour (Kornerup and Wanscher 1978).
From the leaf litter. Asexual morph Synnemata produced in the form of aggregated mass of parallel bundles hyphae, hyaline. Vegetative hyphae aggregated in loose parallel bundles, rope like, branched to unbranched, anastomoses, smooth walled, thin and thick, hyaline, up to 3μm wide. Setae and hyphopodia absent. Chlamydospores absent. Conidiophores arising from superficial hyphae, forming sporodochia, unbranched to branched, branching unilateral to bilateral verticillate, hyaline, smooth walled, 9.68–127.7 × 1.97–2.77 μm. Conidiogenous cells polyblastic and polyphialidic, hyaline, smooth walled, 4.71–35.91 x 1.35–3.46 μm. Phialides solitary or in group of 2–4, sometimes directly produced from superficial hyphae, smooth walled, cylindrical, attenuated, metulae 1–2, primary metulae solitary, spatulate, variable in size (8.59–10.27 ×1.05–1.83 μm), secondary metulae paired (6.98 × 2.12 μm), 8.29–25.36 × 1.35–3.58 μm. Conidia produced in long chain from polyblastic and polyphialidic conidiogenous cell, sometimes conidia accumulated in the form of chevron or zipper like, produced in loose mass on the tip of phialides, oval to broadly ovoid, allantoid to fusoid, smooth walled, base broader, sometimes base rarely tapered, base truncate, tip obtuse, aseptate, hilum unthickened, 1–2 gutulate, smooth walled, hyaline, 2.22– 6.43 × 1.20 – 4.00 μm ( = 3.84 × 2.17 μm, n=30). Sexual morph: Undetermined.
Culture characteristics: on semi-synthetic agar medium PDA white (6A1) to orange white (6A2), reaching 3.0 cm diam. in 5 days at 25°C, forming synnemata with filamentous margin, reverse wrinkled, pale orange (5A3); on PCA (Potato Carrot Agar) white (6A1), reverse yellowish white (4A2), reaching 3.7 cm diam. in 5 days at 25°C, forming dome like structure, pale orange (5A3) exudate. No sporulation on PDA and PCA but abundantly sporulating on grass leaf agar.
Materials examined: INDIA, Maharashtra, Pune District, from leaf litter, 1October 2020, K.S. Pawar, NFCCI 5166 (holotype, ex type living culture) (National Fungal Culture Collection of India- WDCM 932).
Sequence Data: ITS: ON059361; LSU: ON059433
Notes:
Synnemellisia punensis will be a new species from India (Pawar K.S., Singh P. N., Singh S. K., 2022) in this isolate the sexual stage was not found from in vitro- culture on the holotype.
Importance and role: To our knowledge for the first time, this fungus was isolated from leaf litter collected from, District Pune, Maharashtra, India. This fungus was isolated from leaf litter; therefore, it can play role as a litter decomposition.
Importance of genus to humans or ecosystem: Not studied
Industrial relevance and applications: Not studied
Quarantine significance: Synnemellisia punensis was isolated from litter, as Asexual morph. In such condition it may not disassociate from litter and thus it may not provide a pathway for this fungus. Therefore, quarantine assessment is probably not required unless it proves to be nonpathogenic to plant and livestock (Quarantine significance).
Biochemical importance of the species: not studied.
Chemical diversity or applications: not studied
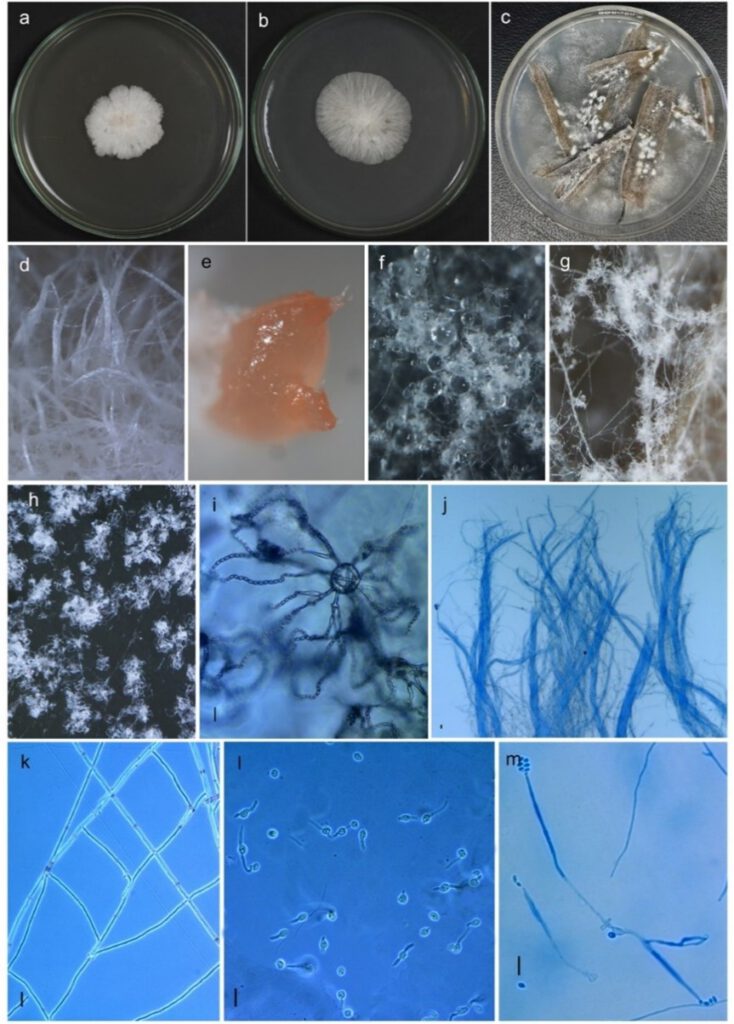
Fig 1 Synnemellisia punensis NFCCI 5166 (holotype). a–b Colony morphology on PDA (front view). c colony morphology on grass leaf media. dStereomicroscopic surface view of colony grown on PDA with hyphal bundles forming synnemata. e Stereomicroscopic surface view of colony grown on PCA with pale orange exudate. f Stereomicroscopic surface view of colony grown on grass leaf media with watery exudate. g–h Whitish puffy mass of sporodochia. i Anastomoses with phialides bearing long chains of conidia. J parallel hyphal bundles of hyphae forming synnemata. k anastomosed hyphae. l Bipolar germination in conidia. m Germinated conidia directly producing phialide with conidia. Scale bars i–m = 10 μm
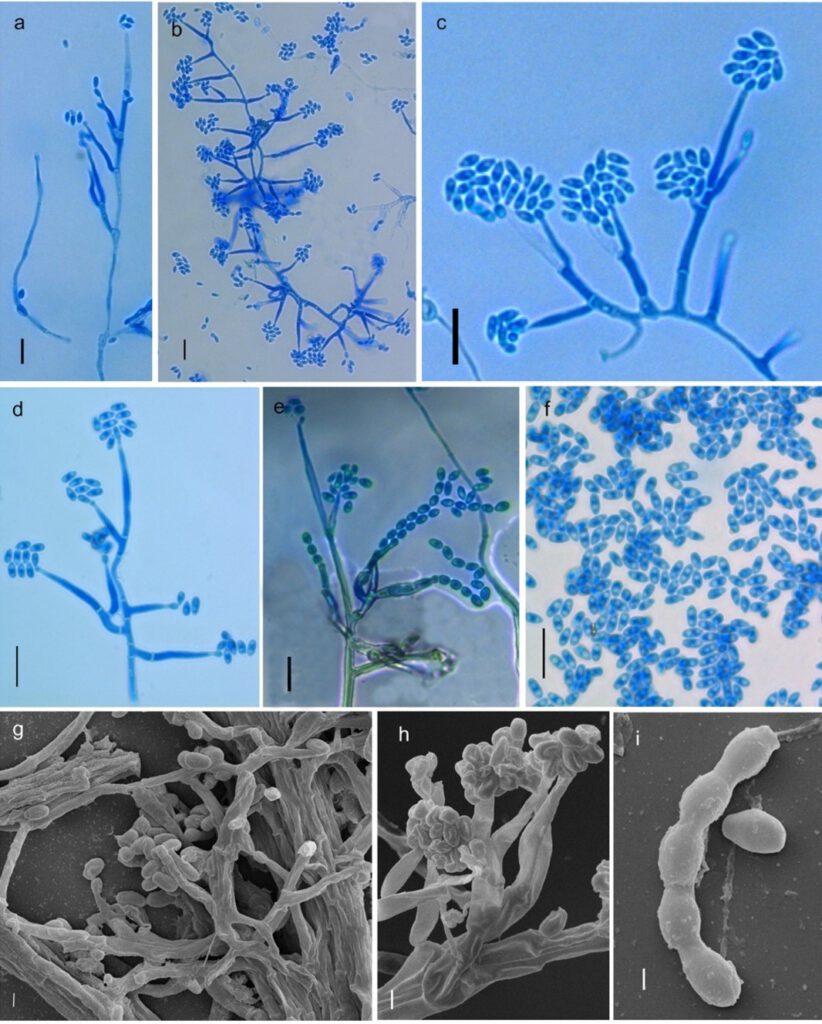
Fig 2 Synnemellisia punensis NFCCI 5166 (holotype). a–b Conidiophores with phialides bearing conidia. c bilateral growth of phialides. c– Conidiophores with phialides bearing conidia (in higher magnification). d Conidiophores with phialides bearing Chevron/ zipper like arrangement of conidia arrangement of spores. e Conidiophores, phialides with chains of conidia. f Numerous conidia. g SEM images of branched conidiophores, phialides & conidia. h SEM images of Conidiophore primary & secondary metulae bearing phialides with conidia. i SEM images of chain of conidia. Scale bars a–f = 10 μm, g–h = 2 μm, i = 1 μm
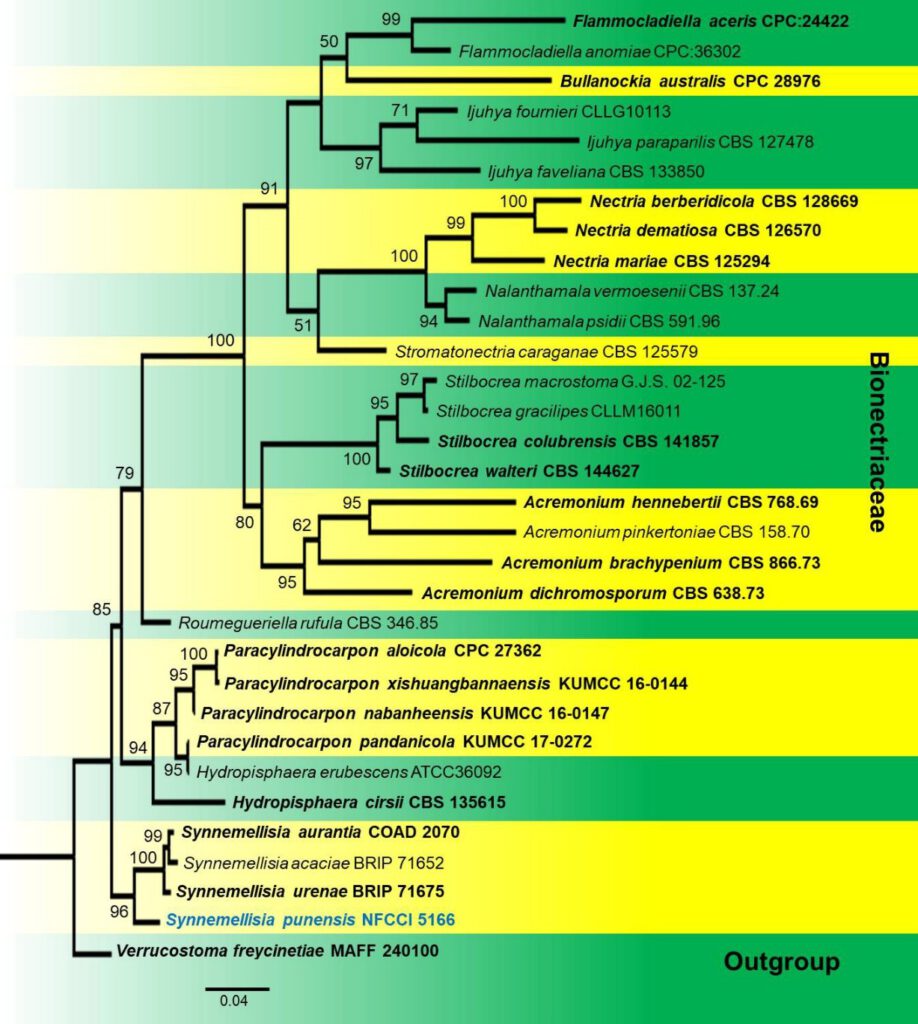
Fig. 3 Phylogenetic tree of Synnemellisia punensis (NFCCI 5166) inferred from Maximum-Likelihood analysis based on combined sequence data of ITS, LSU. Verrucostoma freycinetiae MAFF 240100 was used as outgroup. The analysis involved 32 nucleotide sequences. Evolutionary analyses were conducted in IQ–TREE multicore version 1.6.11 (Nguyen et al., 2015) by the Maximum–Likelihood method using the best suitable model (TNe+I+G4 model). Ex-type strains are in bold and newly generated sequence is in blue. One–thousand bootstrap replicates were analyzed to get ultrafast bootstrap values, and the values above 50% were represented on nodes in the tree.
