Nigrograna kunmingensis T.Y. Du & Tibpromma, sp. nov.
MycoBank number: MB 559865; Index Fungorum number: IF 559865, Facesofungi number: FoF 12956; Fig.X
Etymology: named after the city Kunming from which the holotype was collected.
Holotype: XXXXX
Saprobic on dead stems of Gleditsia sinensis. Sexual morph: Ascomata 300–500 µm high × 390–450 µm diam. (x̅ = 356 × 413 µm, n = 10), aggregated in a disc, immersed in substrate, appearing as black irregular protrusions and cracks, subglobose to globose, brown to dark brown, hairs of ascomata 2–3.5 wide, brown, septate, branched. Ostiole inconspicuous, without papillate. Peridium 20–60 µm wide, comprising several layers, thick-walled cells, comprising brown to dark brown cells of textura angularis. Hamathecium comprising 1.5–2.5 µm wide, filiform, hyaline, septate, branched pseudoparaphyses. Asci (56–)65–80 × 10–12.5(–14) µm (x̅ = 70.3 × 11.4 µm, n = 20), bitunicate, fissitunicate, 8-spored, cylindrical to clavate, straight or slightly curved, with a short pedicel, apically rounded. Ascospores 13–16 × 5–6(–7) µm (x̅ = 14 × 5.4 µm, n=30), uniseriate, brown to dark brown, broadly fusiform or inequilateral, with slightly obtuse ends, upper part or second cell slightly wider, 3-septate when mature, slightly constricted at the septum, straight or slightly curved, guttulate, without appendages. Asexual morph: Undetermined.
Culture characteristics: Ascospores germinated on PDA within 24 h at 28 ℃ and germ tubes were produced from several cells. Mycelium raised, entire, white aerial hyphae; yellow to brown in reverse.
Material examined: CHINA, Yunnan Province, Kunming City, Kunming Institute of Botany, on dead stems of Gleditsia sinensis, 24 March 2021, S.C. Karunarathna, (KMD8), (XXXXXX, holotype); extype culture XXXXXX, XXXXXX.
GenBank numbers: LSU: XXXXXX, ITS: XXXXXX, SSU: XXXXXX, TEF1: XXXXXX.
Notes: In the present phylogenetic analyses, Nigrograna kunmingensis clusters with the sister taxon N. magnoliae (MFLUCC 20-0020, MFLUCC 20-0021) with 100% ML and 1.00 PP statistical support (Fig. X). Nigrograna kunmingensis and N. magnoliae are similar in the shape and size of asci and ascospores. However, N. kunmingensis differs from N. magnoliae in having ascomata with hairs, aggregated in a disc, immersed in the substrate, septate and branched pseudoparaphyses, and uniseriate, brown to dark brown, broadly fusiform or inequilateral ascospores, while N. magnoliae has immersed to erumpent ascomata, septate pseudoparaphyses, and uni to bi-seriate, yellowish-brown to brown, ellipsoid ascospores (Wanasinghe et al. 2020). Therefore, based on both phylogenetic analyses and morphological comparison, N. kunmingensis associated with Gleditsia sinensis is introduced as a new species from China .
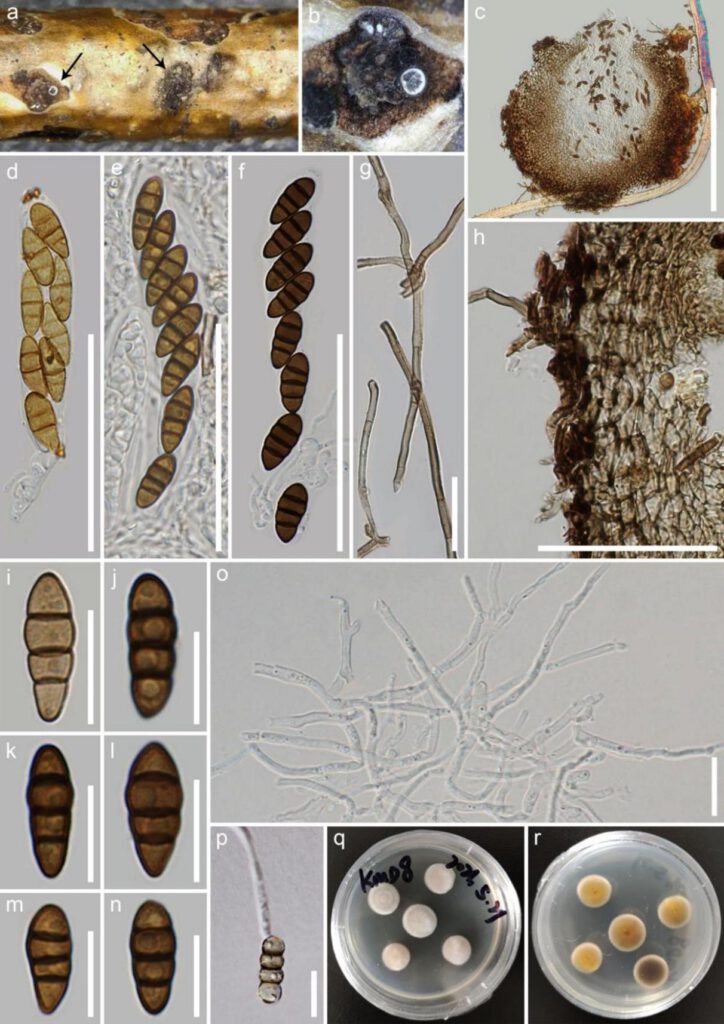
Fig. X Nigrograna kunmingensis (XXXXXX, holotype) a, b Appearance of ascomata on the substrate. c A section through ascoma. d-f Asci. g Hairs of an ascoma. h Peridium. i-n Ascospores. o Pseudoparaphyses. p Germinating ascospore. q, r Colony on PDA from above and below. Scale bars: c=200 µm, d-f=50 µm, g=20 µm, h=50 µm, i-p=10 µm.
