Neomycoleptodiscus alishanense Tennakoon, C.H. Kuo & K.D. Hyde, sp. nov.
MycoBank number: MB; Index Fungorum number: IF; Facesoffungi number: FoF 10790; FIGURE XX
Etymology: The epithet “alishanense” refers to the locality mountain (Ali Shan), from where the taxon was collected.
Holotype: MFLU 19-1732
Saprobic on senescent leaves of Trachycarpus fortune (Hook.) H.Wendl. (Arecaceae). Sexual morph: Undetermined. Asexual morph: Coelomycetous. Conidiomata sporodochial, 120–150 µm high, 130–160 µm diam. ( = 135 × 145 μm, n = 30), superficial, dark brown to black, circular, dull, undulate, umbonate, rough. Conidiogenous cells evanescent. Conidia 16–19 × 3–4 µm ( = 17.5 × 3.6 μm, n = 30), 1-septate, cylindrical, hyaline, guttulate, straight or slightly curved, with filamentous appendages at both ends, 7–7.6 μm long, 0.4–1 μm wide, smooth walled.
Culture characteristics: Colonies on PDA reaching 15 mm diameter after 2 weeks at 25 °C, colonies medium dense, circular, convex, surface slightly smooth with entire edge, effuse, velvety to hairy, margin well-defined and curled, colony from above: grey at the margin, brown at the centre; reverse, light brown to grey at the margin, brown to black at the centre; mycelium grey to greenish with tufting; not producing pigments in PDA.
Material examined: Taiwan, Chiayi, Fanlu Township area, Dahu forest, dead leaves of Trachycarpus fortune (Arecaceae), 06 August 2019, D.S. Tennakoon, GSP014A (MFLU 19-2732, holotype), ex-type living culture, MFLUCC 19-0390; ibid. 11 August 2019, GSP014B (NCYU 19-0006, paratype), NCYUCC 19-0391; ibid. 13 August 2019, GSP014C (NCYU 19-0401), NCYUCC 19-0392.
GenBank numbers: MFLUCC 19-0390: LSU= ON024150; ITS: ON024153. NCYUCC 19-0391: LSU: ON024151; ITS: ON024154. NCYUCC 19-0392: LSU: ON024152; ITS: ON024155
Notes: The morphological characteristics of our collection (MFLU 19-2732 and NCYU 19-0006) tally with the type of Neomycoleptodiscus in having superficial, dark brown to black, circular, sporodochial conidiomata and 1-septate, cylindrical, hyaline, straight or slightly curved conidia with filamentous appendages at both ends (Hernández-Restrepo et al. 2019). Multi-gene phylogeny (LSU, SSU and ITS) generated herein indicates that our collection forms a strongly supported lineage sister to the clade containing Neomycoleptodiscus venezuelense and Mycoleptodiscus endophyticus (85% ML, 1.00 BYPP, Fig. XX). Our collection can be distinguished from Neomycoleptodiscus venezuelense in having smaller conidia (16–19 × 3–4 µm), whereas N. venezuelense has larger conidia (18–27 × 3–5 µm). In addition, a comparison of the 632 nucleotides across the ITS (+5.8S) gene region of our collection and Neomycoleptodiscus venezuelense (CBS 100519) reveals 21 base pair differences (3.32%). Therefore, based on both morphology and phylogeny evidence, we introduce our collection as a new species, Neomycoleptodiscus alishanense from dead leaves of Trachycarpus fortune (Arecaceae). Mycoleptodiscus endophyticus was introduced by Tibpromma et al. (2018) from healthy leaves of Freycinetia sp. (Pandanaceae) as an endophytic species based on mycelial characteristics. However phylogenetically, Mycoleptodiscus endophyticus has a close relationship with Neomycoleptodiscus species (Hernández-Restrepo et al. 2019). Therefore, further taxonomic work is needed to precisely resolve proper morphology based identification and relationships between Mycoleptodiscus endophyticus and Neomycoleptodiscus species.
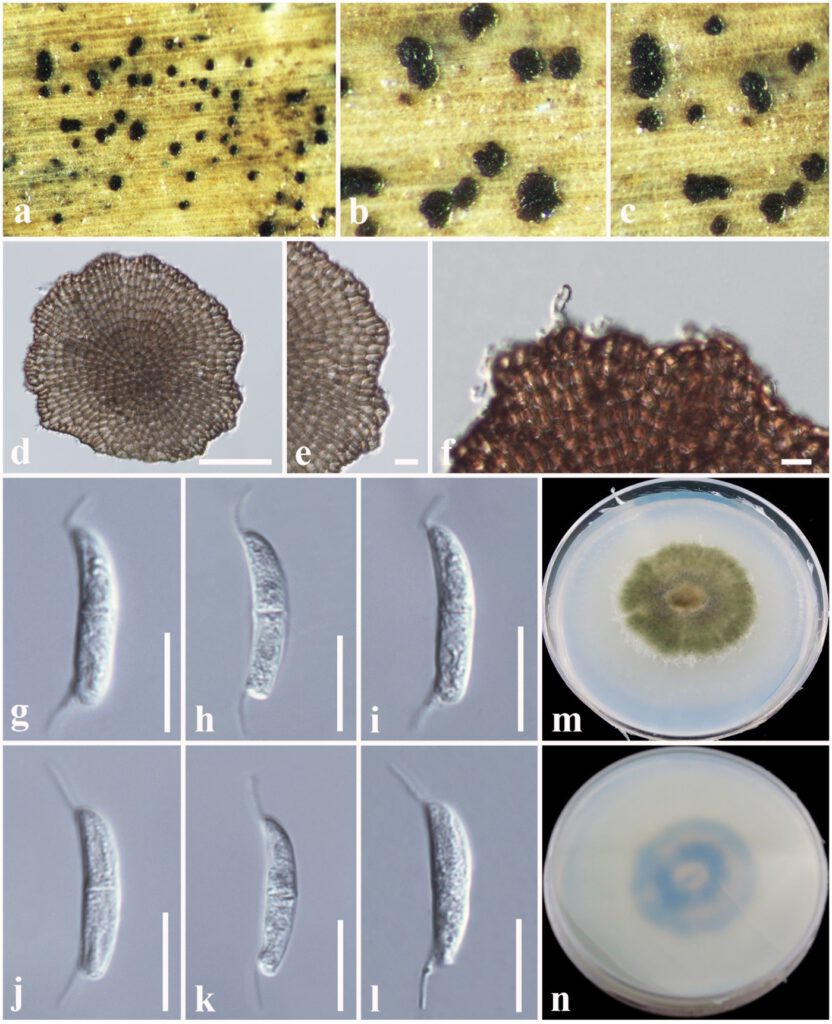
Fig. XX Neomycoleptodiscus alishanense (holotype, MFLU 19-2732) a, b. Appearance of conidiomata on host. c. Close-up of conidiomata. d. Sporodochia. e, f. Conidiomata wall and conidiogenous cells. g–l. Conidia. m. Colony from above. n. Colony from below. Scale bars: d = 50 µm, e, f = 20 µm, g–l = 10 µm.
