Leptosporella elaeidis Konta & K.D. Hyde, in Hyde et al., Mycosphere 11(1): 683 (2020)
Index Fungorum number: IF 556361; MycoBank number: MB 556361; Facesoffungi number: FoF 06026;
Etymology – Named after the host genus, Elaeis.
Holotype – MFLU 19–0669.
Saprobic on rachis and petioles of Elaeis guineensis Jacq. Sexual morph: Ascomata (including neck) 242–600 μm high × 625–1000 μm diameter (x̅ = 444 × 787 μm, n = 10), solitary, superficial, comprising black, carbonaceous, dome-shaped, raised blister-like areas, subglobose, flattened at the base, ostiolate. Ostiole central, carbonaceous, black, with periphyses. Peridium 63–113 μm wide ( x = 83 μm, n = 10), outer cells merging with the host epidermal cells, composed of dark brown to black cells of textura angularis. Paraphyses numerous, 2–7 µm diameter (x̅ = 4 μm, n = 10), branched, septate, longer than asci. Asci 125–210 × 9–14 μm (x̅ = 163 × 11 μm, n = 20), 8-spored, unitunicate, cylindrical, long-pedicellate, with a J-, wedge-shaped, subapical ring. Ascospores 124– 136 × 2–4 μm (x̅ = 128 × 3 μm, n = 20), fasciculate, yellowish in mass, spiral, filiform, straight or curved, C-shaped or sigmoid, aseptate, rounded at the apex, pointed at the base, without polar mucilaginous appendages, smooth-walled. Asexual morph: Undetermined.
Material examined – THAILAND, Suratthani Province, on dead rachis and petioles of Elaeis guineensis Jacq. (Arecaceae), 21 July 2017, Sirinapa Konta SRWD14b, MFLU 19-0669, holotype.
GenBank numbers – ITS: MK659767, LSU: MK659772, SSU: MK659774, tef1: MN883560.
Notes – The ascospores of Leptosporella elaeidis failed to germinate, and therefore the DNA was obtained directly from fruiting bodies. Phylogenetic analysis placed our collection with L. arengae, 61% ML (Fig. 8). Leptosporella elaeidis differs from L. arengae in having larger ascomata, asci and ascospores and lack of polar mucilaginous appendages. This is the first record of L. elaeidis on Elaeis guineensis and the fourth record of Leptosporella species from a palm tree (Arecaceae).
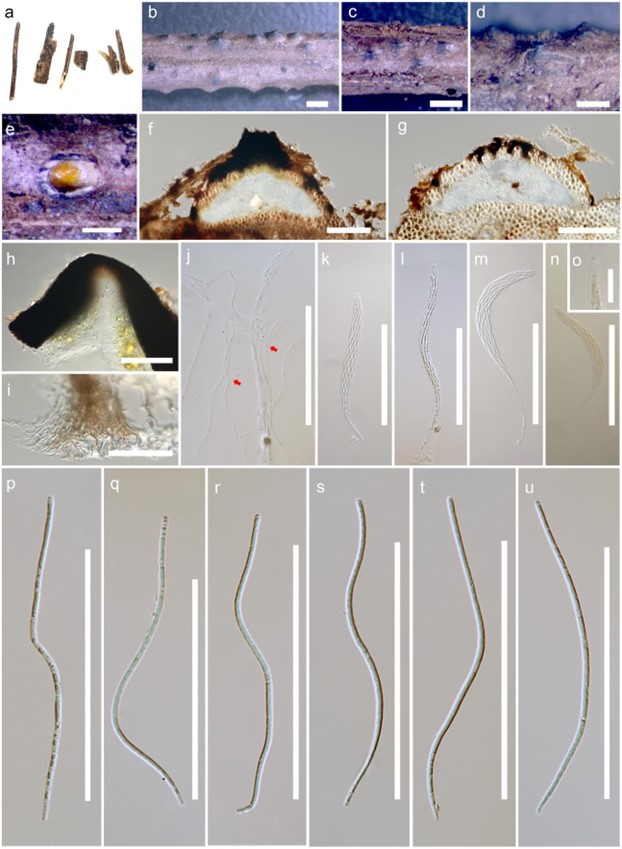
Figure 148 – Leptosporella elaeidis (MFLU 19-0669, holotype). a, b Appearance of ascomata on host substrate. c, d Close up of ascomata. e Yellowish ascospore mass. f, g Sections of ascoma. h Neck with periphyses. i Peridium. j Paraphyses (branch and septate at red arrow). k-m Asci. n-o J-, reaction of apical ring. p-u Ascospores. Scale bars: b, c = 1000 μm, d, e = 500 μm, f, g = 200 μm, h, j-n, p-u = 100 μm, i = 50 μm, o = 20 μm.

Figure 8 – Phylogram generated from maximum likelihood analysis based on combined LSU and ITS sequence data of Chaetosphaeriales and Tracyllalales taxa. Ninety-six strains are included in the combined analyses which comprised 1695 characters (1081 characters for LSU, 614 characters for ITS) after alignment. Neurospora crassa MUCL 19026 and Gelasinospora tetrasperma CBS 178.33 (Sordariaceae, Sordariales) are used as outgroup taxa. Single gene analyses were carried out and the phylogenies were similar in topology and clade stability. Tree topology of the maximum likelihood analysis is similar to the Bayesian analysis. The best RaxML tree with a final likelihood value of – 23777.689886 is presented. Estimated base frequencies were as follows: A = 0.231060, C = 0.264793, G = 0.308265, T = 0.195882; substitution rates AC = 1.388486, AG = 1.836207, AT = 1.649563, CG = 0.971659, CT = 6.316962, GT = 1.000000; gamma distribution shape parameter a = 0.460297. Bootstrap support values for ML greater than 75% and Bayesian posterior probabilities greater than 0.95 are given near the nodes. Ex-type strains are in bold. The newly generated sequences are indicated in blue.
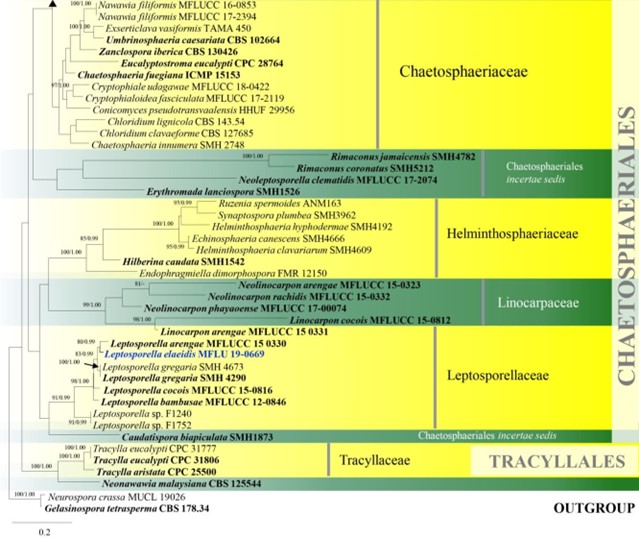

Figure 8 – Continued.
