Bullimycetaceae K.D. Hyde & Hongsanan, fam. nov.
MycoBank number: MB 557986; Index Fungorum number: IF 557986; Facesoffungi number: FoF 09416;
Type genus: Bullimyces Ferrer et al.
Saprobic on submerged, dead woody debris. Sexual morph: Ascomata on submerged wood immersed to erumpent, globose to subglobose, membranous, black, ostiolate. Neck cylindrical, dark brown. Paraphyses thin‑walled globose, hyaline cells, filling the centrum, deliquescent. Asci 8‑spored, unitunicate, cylindrical, thin‑walled, lacking an apical pore and apical structures, deliquescing early in some species. Ascospores uniseriate, broadly ellipsoidal‑fusiform to ellipsoidal, hyaline becoming dark yellow with age, septate, not constricted at septa, thick‑walled, with or without a gelatinous sheath and appendages. Asexual morph: Undetermined.
Notes: Bullimyces was introduced by Ferrer et al. (2012) with three species from freshwater habitats in Costa Rica. Previous phylogenetic studies placed this genus in Sordariomycetidae genera incertae sedis (Ferrer et al. 2012; Maharachchikumbura et al. 2015). With more genera and families being introduced to Sordariomycetes, phylogenetic analysis of Luo et al. (2019) showed that Bullimyces formed a well-supported clade within Diaporthomycetidae, and they transferred Bullimyces to Diaporthomycetidae genera incertae sedis. Our phylogenetic analysis obtained a similar result with Luo et al. (2019), and taxa of Bullimyces clustered with Ceratolenta, forming a distinct clade within Diaporthomycetidae (Fig. 1). The divergence time for Bullimyces is estimated as 81 MYA (Figs. 2, 3), which falls in the family range (50–130 MYA).
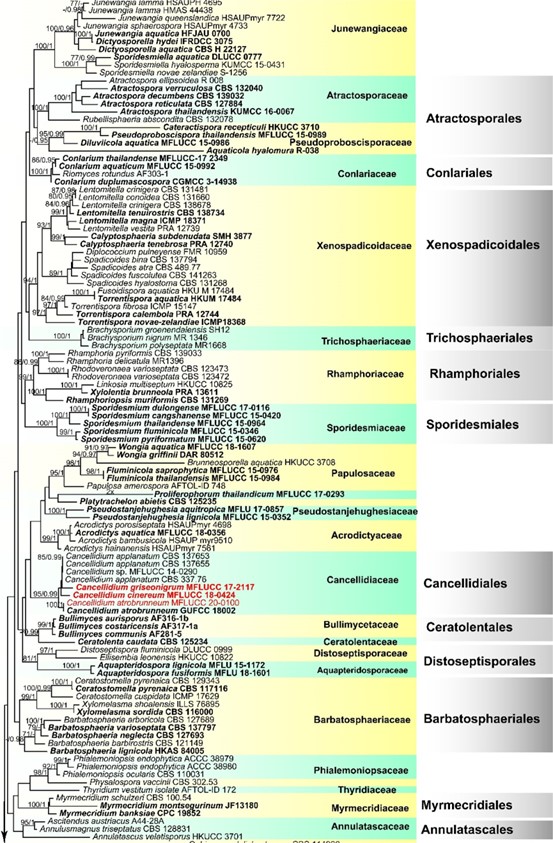
Fig. 1 Phylogenetic tree based on RAxML analyses of combined LSU, SSU, TEF1‑α and RPB2 sequence data. Maximum likelihood bootstrap ≥ 75% (MLBS) and Bayesian posterior probabilities ≥ 0.95 (PP) are indicated at the nodes. The new species are in red and ex‑ type strains are in bold. The tree is rooted with Chlorociboria aeruginosa (AFTOL‑ID 151) and Dermea acerina (AFTOL‑ID 941)
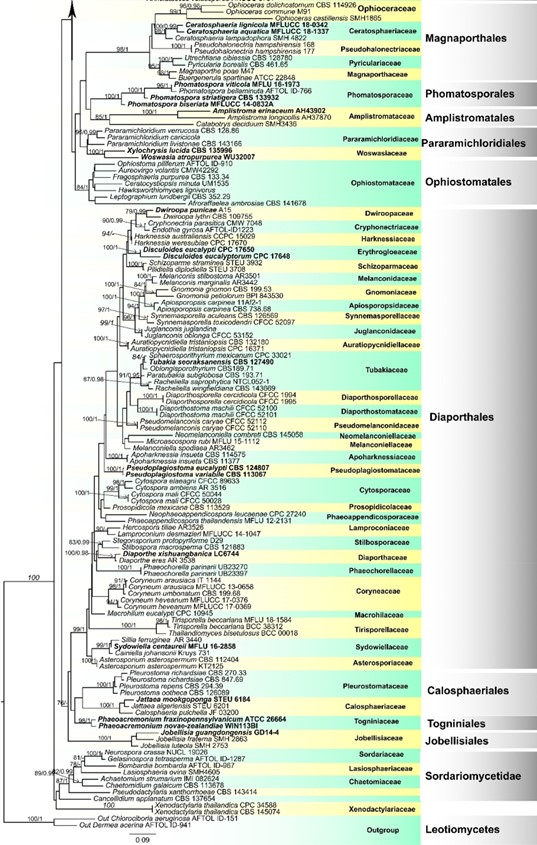
Fig. 1 (continued)
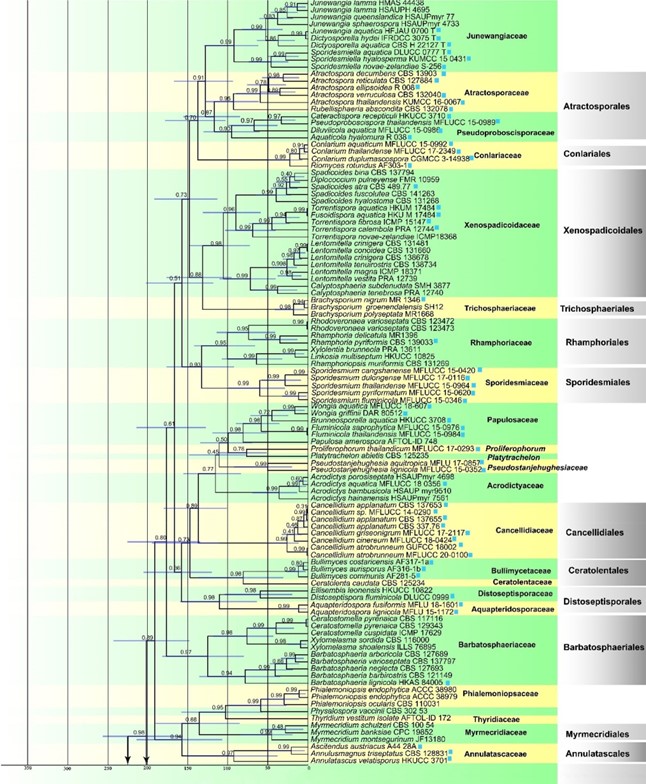
Fig. 2 Maximum clade credibility (MCC) tree with divergence time estimates for Diaporthomycetidae. Numbers at nodes indicate posterior probabilities (pp) for node support; bars correspond to the 95% highest posterior density (HPD) intervals. The blue squares indicate those species from freshwater habitats, numbers in the red circles indicate the fossil (1) and secondary (2) points
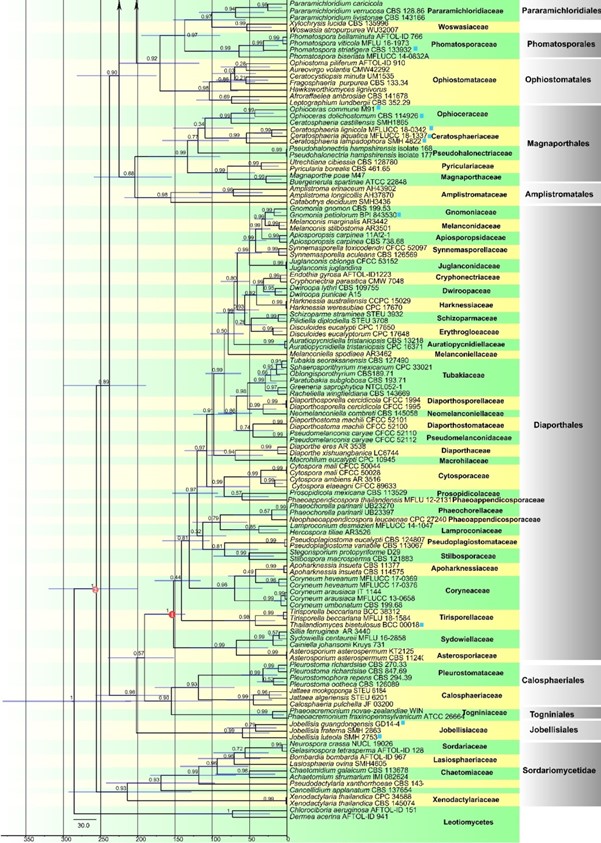
Fig. 2 (continued)
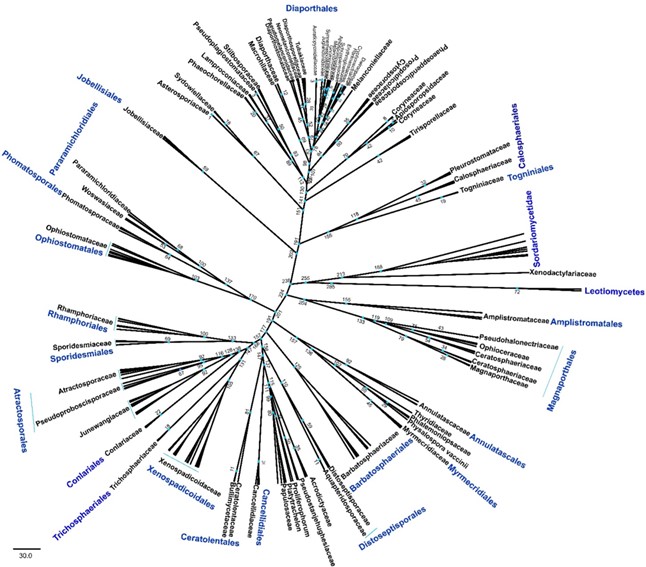
Fig. 3 The accepted families and orders in Diaporthomycetidae with divergence time estimations are shown in the MCC tree
Genus
Bullimyces
