Beltraniella jiangxiensis P. Razaghi, M. Raza & L. Cai
Mycobank number: MB 844849; Index Fungorum number: IF 844849; Facesoffungi number: FoF 12674;
Etymology – The name refers to the location it was collected.
Holotype – HMAS 352097.
Asexual morph: Setae numerous, erect, arising from radially lobed basal cells, straight or flexuous, unbranched, single or in small groups, thick-walled, coarsely verrucose, olivaceous brown, paler towards apex, 81–304 μm long, 5–8.5 μm wide, tapering to a pointed apex, up to 16-septate, arising from a dark brown, swollen, radially lobed basal cell, 11–22 μm diameter. Conidiophores macronematous, short, simple or branched at apical regions, 3–6 septate, reduced to conidiogenous cells, verrucose, thin-walled, swollen at the base, subhyaline to pale olivaceous, arising from basal cells of setae or from separate, 36–65 × 4–7 µm. Conidiogenous cells polyblastic, integrated, determinate, terminal, geniculate, denticulate, cylindrical, oblong, hyaline to subhyaline, smooth, 9–19 × 4–6 μm (x̄ =12.62 ± 3.27 × 4.68 ± 0.59 μm). Separating cells obclavate, thin-walled, smooth, hyaline to subhyaline, 1-denticulate at each end, 8–11 × 3.5–5 μm (x̄ =9.91 ± 0.89 × 3.95 ± 0.47 μm). Conidia arise directly from conidiogenous cells or from separating cells, aggregated, acrogenous, simple, dry, straight, sometimes verrucose, thin-walled turbinate to pyriform, rostrate to pointed at proximal end, truncate at distal end, hyaline to subhyaline without a hyaline transverse band, 20–24 × 5.5–7.5 μm (x̄ =22.12 ± 1.25 × 6.55 ± 0.64 μm). Sexual morph: Undetermined.
Culture characteristics – Colonies on PDA raised with a concave edge, with undulate edge, dense, colony cream from above and reverse, velutinous, sterile, reaching 55–57 mm diam after 7 days at 25 °C; on SNA dome-shaped, with rhizoid edge, white, reaching 53–60 mm diam after 7 days at 25 °C.
Material examined – China, Jiangxi, Ganzhou, Fengshan, on Camellia sinensis, Aug. 2013, Y. Zhang (HMAS 352097, holotype), ex-type living culture CGMCC 3.23486 = LC3449.
Other specimens examined – China, Jiangxi, Ganzhou, on leaves of Camellia sinensis (Theaceae), Aug. 2013, Y. Zhang, living culture LC15868.
Notes – Two isolates of Beltraniella jiangxiensis formed a well-supported and distinct clade on the ML tree generated from ITS and LSU sequences, closely related to Belt. pandanicola, Belt. portoricensis and Belt. fertilis. Morphologically, our novel species is distinguished from the most closely related species, Belt. pandanicola isolated from Pandanus sp. in Thailand (Tibpromma et al. 2018) by producing more septate (16 μm vs. 2–4 μm) and relatively longer seta (81–304 μm vs. 114–200 μm), longer conidiophores (36–65 μm vs. 20–42 µm) and longer conidiogenous cells (9–19 μm vs. 6–10 μm).
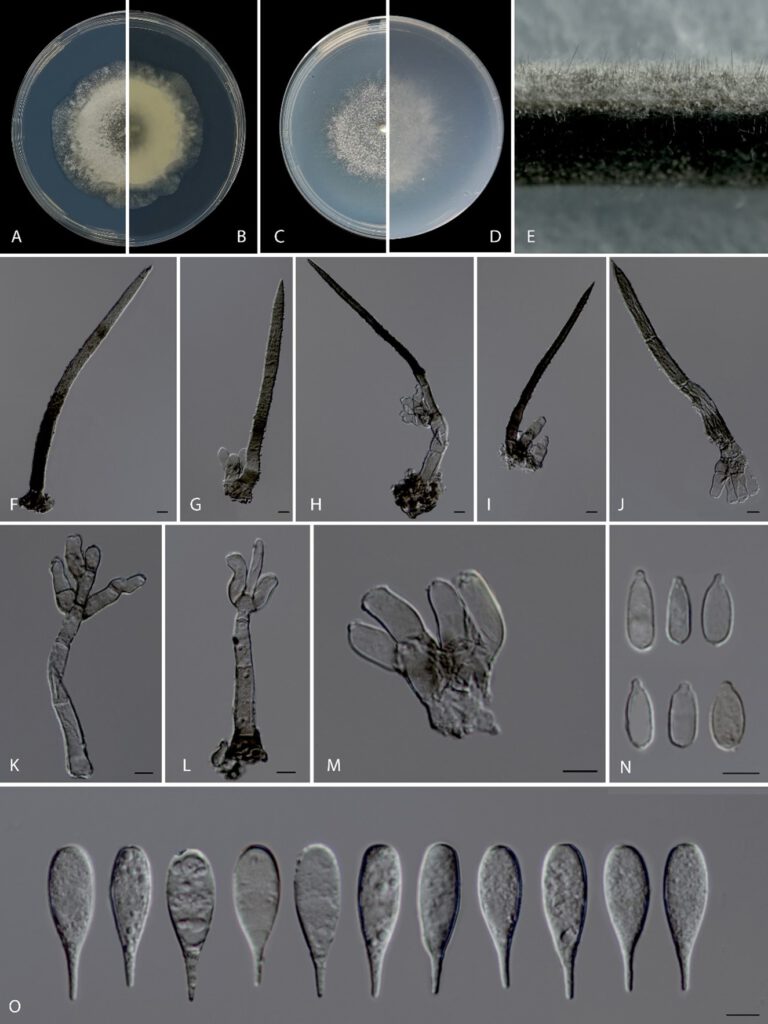
Fig. 1 Beltraniella jiangxiensis (HMAS 352097, holotype) a–d Upper and reverse views of cultures on PDA and SNA after 1 week of inoculation, respectively. e Conidiophores and setae on pine needle on SNA. f–j Setae, conidiophores and conidiogenous cells. k–m Conidiophores and conidiogenous cells. n Separating cells. o Conidia. Scale bars: 5 µm
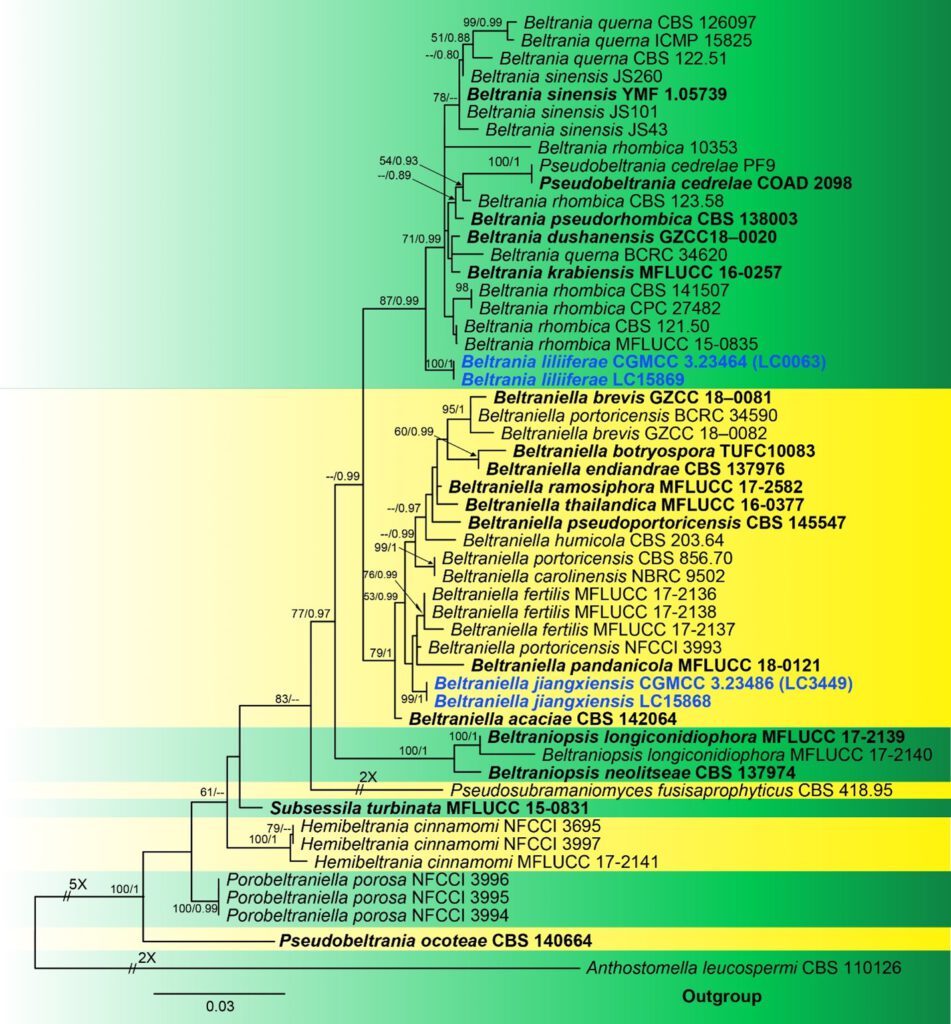
Fig. ****** Phylogenetic tree generated from Maximum likelihood analysis (RAxML) based on combined ITS and LSU sequence data of Beltraniaceae.
